Accelerating Development of a New Single-Molecule Localization and Tracking Technique
By Maged F. Serag, King Abdullah University of Science and Technology
Pioneered more than 30 years ago, single-molecule localization and tracking (SMLT) is a technique for characterizing the motion of individual molecules. By measuring diffusion coefficients and characterizing molecular motion as random, directed, or constrained, scientists can investigate subcellular dynamics in live cells, including viral infection, gene transcription, and the behavior of receptors on cell surfaces.
Despite its relatively long history and many applications, SMLT has several drawbacks. It does not, for example, tell us the molecule’s shape and size or how these change over time. In addition, SMLT is inefficient and sometimes fails to work due to statistical errors resulting from out-of-focus motion of the molecules.
My research group at King Abdullah University of Science and Technology (KAUST) developed a method for measuring single-molecule diffusion that has none of these limitations. Rather than quantifying diffusion from the spatial and temporal components of the molecule’s trajectory, as in traditional SMLT, our MATLAB® based method quantifies diffusion by analyzing the increase of the cumulative area (CA) occupied by the molecule in space over time (Figure 1). We validated our approach with MATLAB by comparing the statistical distribution of diffusion coefficients calculated by traditional SMLT techniques and those calculated with the new CA method. The CA method outperformed traditional SMLT in the reproducible measurement of diffusion dynamics for the DNA molecules we tested.
The core of our work— image processing and performing fitting and mathematical calculations on microscope images—is done using MATLAB. MATLAB offers three key advantages that make it a good fit for our research. First, it is easy to learn. Even though my background is in pharmacy, not programming, I mastered MATLAB well enough to conduct this research in just one month. It would have taken me six times longer to reach a similar level of mastery in a language like C++ or Java®. Second, KAUST has a Total Academic Headcount (TAH) license, which makes it easy for researchers across KAUST to access MATLAB and the large collection of capabilities and functions in its add-on toolboxes anywhere on campus. Third, the SMLT and CA methods are computationally intensive, requiring hundreds of thousands of Gaussian fittings for a single experiment. Parallel Computing Toolbox™ and MATLAB Parallel Server™ enabled me to accelerate these methods and shorten processing times for multiple experiments from days to hours (see sidebar).
Creating Image Sequences of Simulated Particles, Nanospheres, and DNA Molecules
Both SMLT and CA methods involve analyzing a sequence of image frames, typically captured from a microscope, with one or several molecules visible in each frame. We applied the CA method to characterize the motion of particles and calculate diffusion coefficients in three separate scenarios. The first uses simulated data to create the sequence of images. The second and third use sequences of images obtained using a custom-built wide-field epifluorescence microscope in our lab.
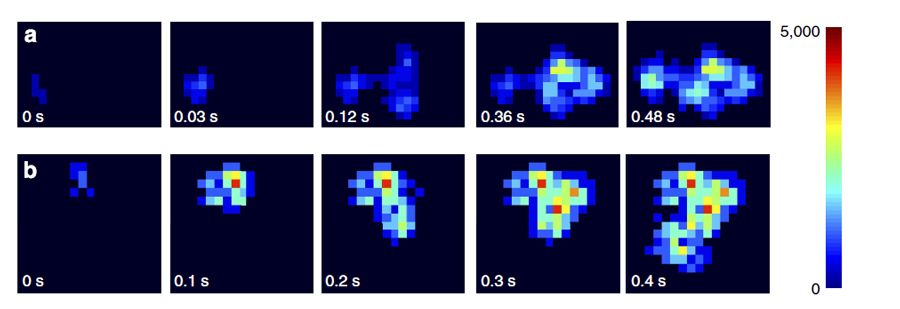
We designed the first scenario to validate the CA method. In MATLAB, we generated random-walk trajectories of particles in 2D space using predetermined diffusion coefficients of 1.0, 1.5, and 2.0 micrometers2/sec. For each step on the random walk, the x and y positions of a particle were used to define the center of a five-pixel cross in a single frame in the image sequence (Figure 2). We then used the CA method to calculate the diffusion coefficient from the simulated particles, and verified that the results (1.10, 1.51, and 1.98 micrometers2/sec, respectively) were in agreement with our predetermined values.
For the second and third scenarios, we tracked yellow fluorescent polymer nanospheres about 0.2 micrometers in diameter and double-stranded DNA molecules of different lengths and topological forms. We captured images of the nanospheres and molecules at a rate of 1 frame per 6.4 ms. We processed these images using both SMLT and CA methods.
Implementing the CA Method
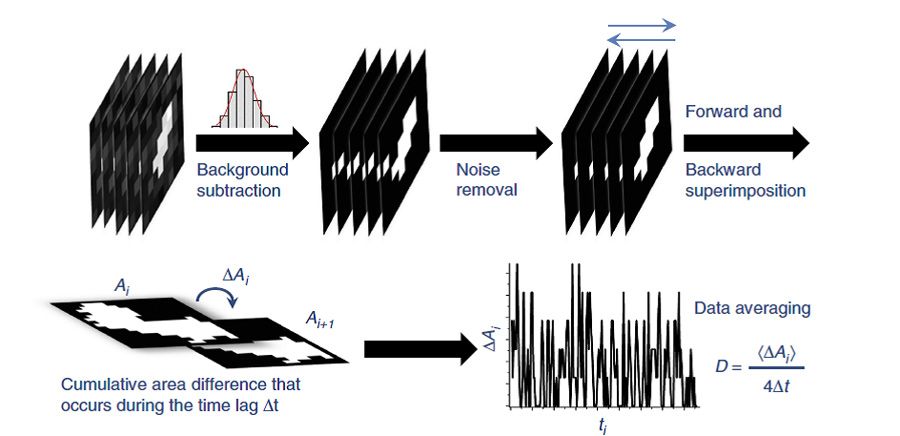
Working in MATLAB, we developed an algorithm to implement the CA method (Figure 3). Using the sequences of thousands of 512 x 512 pixel frames generated through simulation or captured in the lab, the algorithm first invokes Image Processing Toolbox™ functions to remove the background based on an initial threshold. The algorithm calculates this threshold by fitting the frequency distribution of the intensity of all pixels in the frame with a Gaussian function using Curve Fitting Toolbox™.
After removing noise pixels from the frame, the algorithm gradually increases the background threshold until just five pixels remain, defining the area of the space occupied by the molecule in that frame.
When all frames in the sequence have been processed, the algorithm superimposes them to generate the cumulative area occupied by the molecule in each frame and then subtracts the cumulative area of adjacent frames to find the cumulative area difference, which is used to calculate the diffusion coefficient.
Accelerating the Process with Parallel and Distributed Computing
With a single experiment requiring about 200,000 Gaussian fittings, we soon found that running experiments on a single processor took too long to be practical. To shorten processing times we used Parallel Computing Toolbox to perform the computations on a workstation with multiple cores. Using four cores experiments took about three hours, and with 16 cores, just 45 to 50 minutes.
Of course, we often need to run many simulations and experiments to obtain valid statistical results. To further accelerate the process we began running our jobs on 512 cores at a time on the IT Research Computing clusters at KAUST with MATLAB Parallel Server. These clusters offer more than 10000 cores to the users. Using this setup we can complete a set of experiments that took 24 hours on a multicore machine in just 15 minutes.
Visualizing and Interpreting Results
We are currently interpreting the results of our simulations and experiments. With MATLAB we visualize experimental results to better understand how the CA-method is performing compared with the SMLT.
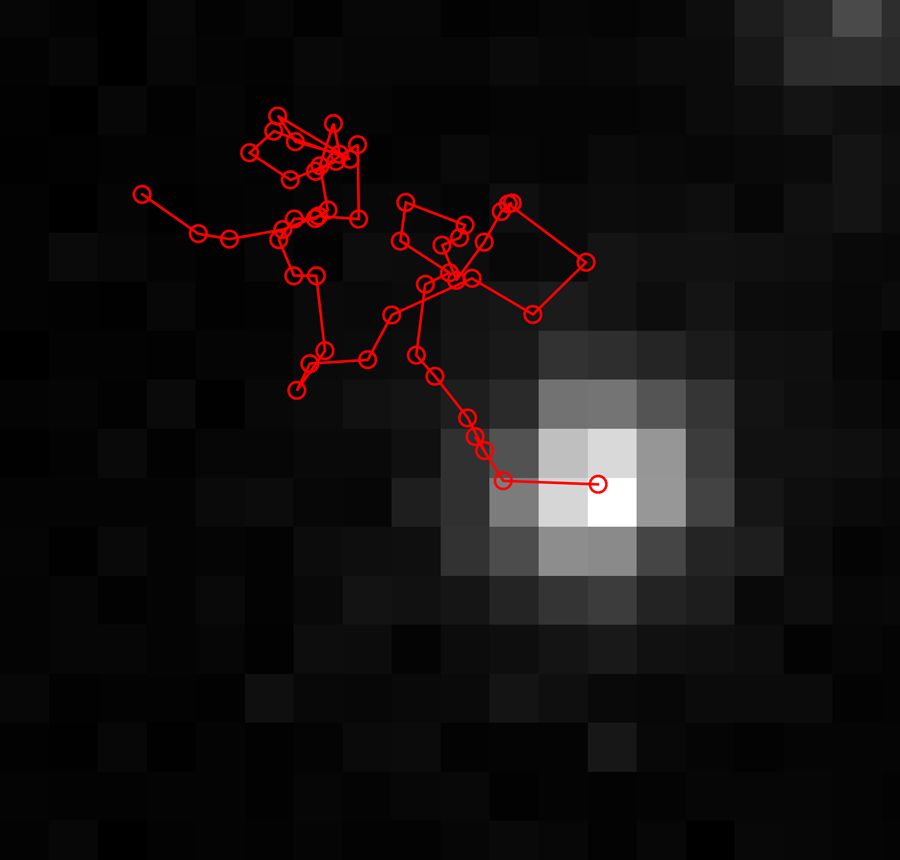
To enable a comparison of the CA-method with traditional SMLT on the same experimental data, we implemented SMLT in MATLAB. Our SMLT algorithm applies 2D Gaussian fittings over the pixels in each frame to determine the position of the molecule’s center of mass. After repeating this process for each frame, the algorithm connects the centers of mass across frames to create a trajectory and then performs mean squared displacement analysis of the trajectories to characterize the molecule’s motion (Figure 4).
We are using dynamic time warping (DTW) techniques implemented in MATLAB to measure similarities and differences between the SMLT and CA-method results. Early results suggest that the CA-method has a smaller statistical error, in addition to the ability to provide scientists with information on molecular size and frequency of conformational changes.
Running MATLAB on Research Computing Clusters
By Dr. Matthijs van Waveren, KAUST IT Research Computing
MATLAB Parallel Server enables researchers at KAUST to run their computationally intensive MATLAB programs on the computer clusters maintained and managed by the university’s IT Research Computing group.
To make it easier for researchers to use the clusters, our group worked with MathWorks® consultants to develop a high-performance computing (HPC) add-on for MATLAB. Researchers can use this add-on from within the MATLAB environment to execute their scripts on hundreds of workers. The add-on takes care of transferring data files and scripts to the cluster, running the jobs, and then transferring the results back to the researcher’s MATLAB environment.
The HPC add-on made it easier for researchers to use clusters for their MATLAB jobs. As a result, demand for cluster time increased dramatically. To meet this demand, we built a virtual cluster using OpenStack and a set of Linux® workstations. We then updated the HPC add-on so that users could run their jobs either on one of the original clusters or on the new virtual cluster. While not as fast as the original clusters, the virtual cluster is available for researchers who do not want to wait for their jobs to be scheduled on the original clusters during periods of high demand.
The author wishes to thank Raymond Norris and Amine El Helou of MathWorks for assistance in developing the HPC add-on.
Published 2016 - 92970v00