Automating Genotoxicity Assays with Imaging Flow Cytometry and Deep Learning
By Paul Rees, Swansea University
Genotoxicity assays are used to assess the damage to genetic material (DNA and RNA) caused by a chemical agent such as a drug, and are often used to test the safety of candidate drugs. In the past, highly trained lab technicians conducted assays manually, using a microscope to inspect thousands of individual cells to identify the biomarkers of DNA damage: the formation of micronuclei (MN) upon cell division (Figure 1). In addition to being labor-intensive and time-consuming, this approach depended on the subjective judgement of each individual technician.
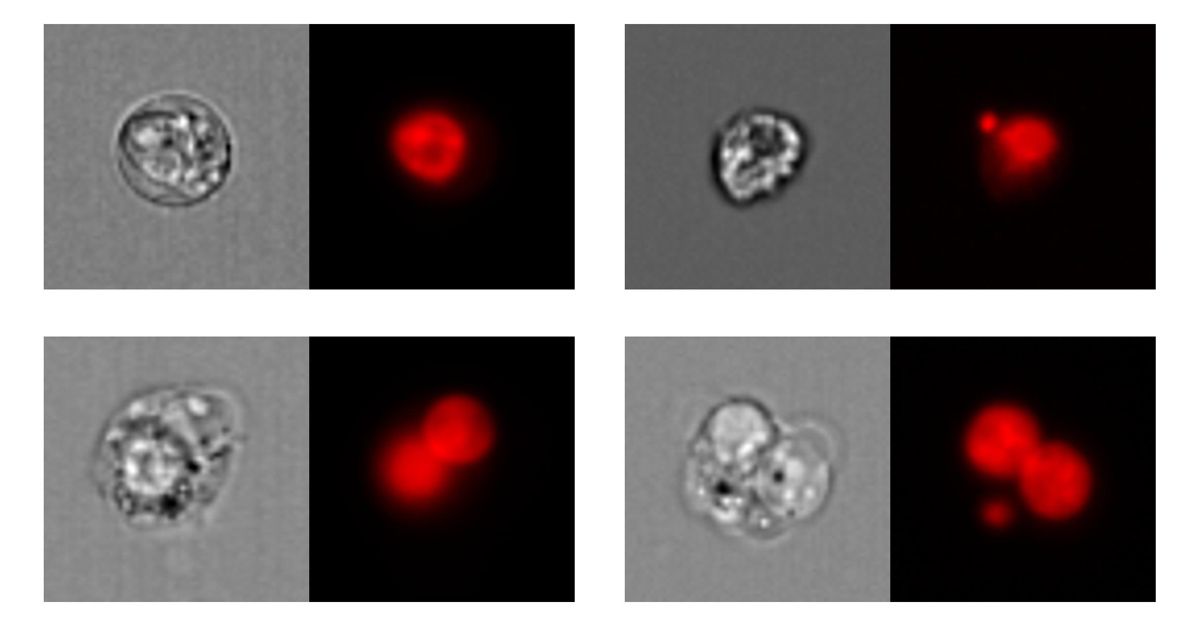
Figure 1. Top left: mononucleated cell; top right: mononucleated cell with micronucleus. Bottom left: binucleated cell; bottom right: binucleated cell with micronucleus. Left: bright-field images; right: nuclear fluorescence images.
My research group at Swansea University has developed an automated approach to genotoxicity and similar studies based on deep learning and imaging flow cytometry (IFC). My collaborator Dr. George Johnson’s lab uses an IFC to collect multichannel image data from individual cells. We use DeepFlow, a deep learning network optimized for use with IFC [1], which enables us to accurately and automatically categorize the images as single nucleus, single nucleus with MN, two nuclei, or two nuclei with MN (Figure 2).
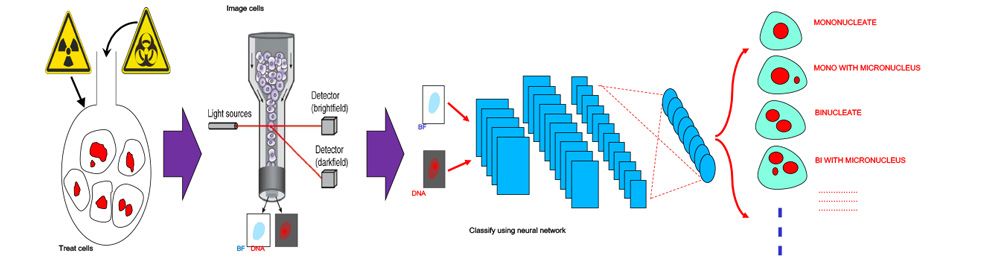
Figure 2. Automated image classification workflow.
This approach eliminates the subjectivity of manual approaches and enables assays to be performed with consistent results in labs around the world. Because we implemented DeepFlow in MATLAB® with Deep Learning Toolbox™, we can send the code to any lab that we collaborate with and know that it will run reliably. Many researchers are already familiar with MATLAB, which means they can easily modify or improve the code and tailor DeepFlow to their specific experimental setups.
Building DeepFlow
We initially implemented DeepFlow using Keras TensorFlow™ but decided to reimplement it in MATLAB so that DeepFlow could be used in virtually any lab in the world. We wanted our software to work no matter what flow cytometry machine a particular lab was using. We didn’t want to worry about dependencies, and we wanted a deep learning network that was easy to understand and to modify.
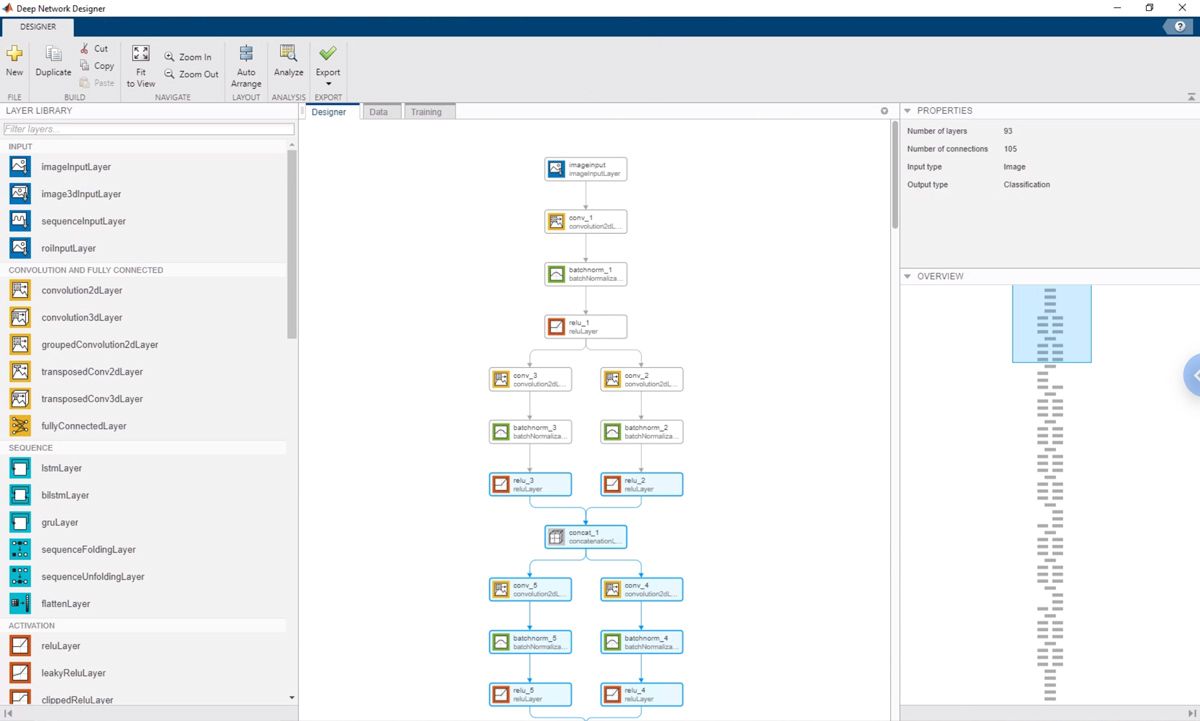
Rather than performing a line-for-line translation of our Keras code, we used the Deep Network Designer app to build, visualize, and train the DeepFlow network. With the Keras code up on one side of the screen and the Deep Network Designer app on the other, we simply replicated the architecture of the initial implementation (Figure 3).
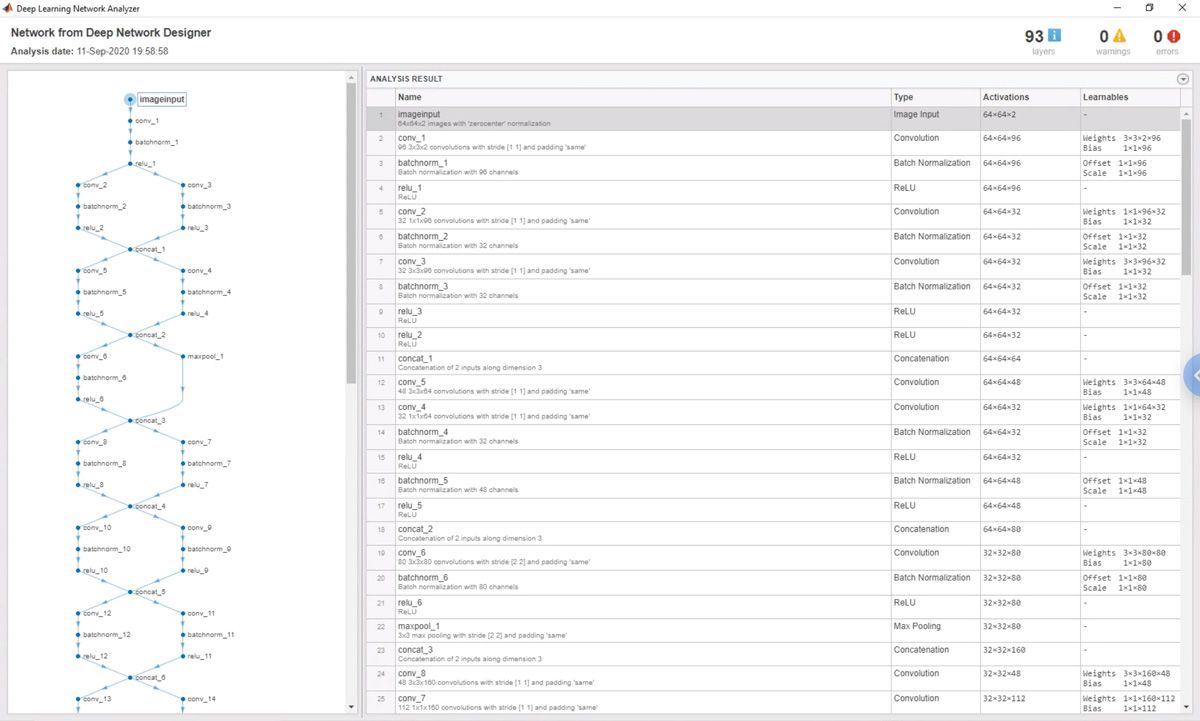
We used the network analyzer in Deep Learning Toolbox to check for errors in the network and its layers (Figure 4). For example, we started with a network designed for images of 200x200 pixels and downsized it to work with the 64x64-pixel images that we get from the IFC, using the network analyzer to verify the image size at each convolutional layer in the network. Our collaborators also use the network analyzer when they make changes to the network with the Deep Network Designer app.
Using DeepFlow in Genotoxicity Assays
In our experimental setup, we use an IFC capable of processing 10,000 cells within minutes. We capture bright-field images as well as fluorescence images of nuclei and micronuclei stained with a solution that makes the DNA more visible (Figure 5).
![Figure 5. CNN for classifying and visualizing IFC data. [1]](https://ww2.mathworks.cn/en/company/technical-articles/automating-genotoxicity-assays-with-imaging-flow-cytometry-and-deep-learning/_jcr_content/mainParsys/image_0_copy_copy_co_452274873.adapt.full.medium.jpg/1709787972353.jpg)
Figure 5. CNN for classifying and visualizing IFC data. [1]
We bring the IFC data into MATLAB as a MATLAB datastore. We preprocess it with conventional image processing techniques to renormalize each image based on its intensity and to ensure that each image is in focus, with the cell completely in the frame. We use edge detection, for example, to identify clean edges, which indicate the image is in focus, and perfectly flat edges, which indicate that the camera has not captured the entire cell.
We trained the DeepFlow CNN on more than 2000 manually classified images. Once we have a normalized, clean set of data from the IFC, we use the trained network to classify images as having mono-, bi-, tri-, or tetra-nucleated cells with or without MN. Finally, using a well-established formula that calculates the percentage of cells falling into each category, we can assess the toxicity of the agent used to treat the cells.
We’ve found that the layer immediately above the final classification layer in the DeepFlow network is particularly valuable for understanding how the trained CNN is working. To parse the information embedded within this layer, we use MATLAB to apply a t-distributed stochastic neighbor embedding (tSNE) algorithm for visualizing high-dimensional data (Figure 6). These visualizations can reveal nuances in the image data that are barely perceptible in a manual inspection. For example, the relationship between cells classified as binucleate and mononucleate with a micronucleus demonstrates that size determines the difference between a normal nucleus and a micronucleus.

Figure 6. A tSNE visualization of data showing typical cell types from the genotoxicity assay.
Extending DeepFlow Principles to Blood Quality Assessment Using Weak Supervised Learning
In addition to the genotoxicity study, we’ve used deep learning in a variety of analysis and classification applications. Recently, for example, my colleagues and I used a CNN and weakly supervised learning to study the degradation of red blood cells (RBCs) over time [2]. RBCs in blood stored for transfusions develop storage lesions, observed as changes in cell morphology, which are often assessed manually with a microscope. Manual assessment is incredibly time-consuming, and we noticed that different experts often produced different scores.
For the first part of the RBC study, we proceeded as we had with the genotoxicity study, training a CNN with images that had been manually labeled as belonging to one of several possible morphologies, or phenotypes (Figure 7). The trained network achieved greater than 76% agreement with an expert in classifying the morphologies, which is comparable to the approximately 79% agreement seen between experts.
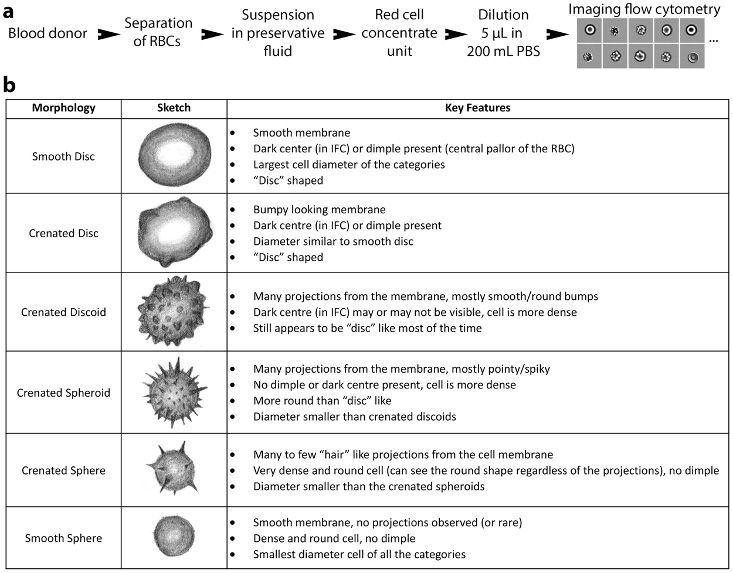
Figure 7. RBC morphologies.
For the second part of the study, we eliminated subjective human labeling and trained a weakly supervised neural network, ResNet50, using only the amount of time the blood had been in storage. When we visualized the results with the tSNE-based technique used in the genotoxicity study, we found that the network had learned to extract single-cell features that revealed a chronological progression of morphological changes (Figure 8). We realized that this progression could be used to predict blood quality and the expiration date of stored blood without human-curated annotation, reducing blood wastage and helping ensure that unsuitable blood is not used in transfusions.
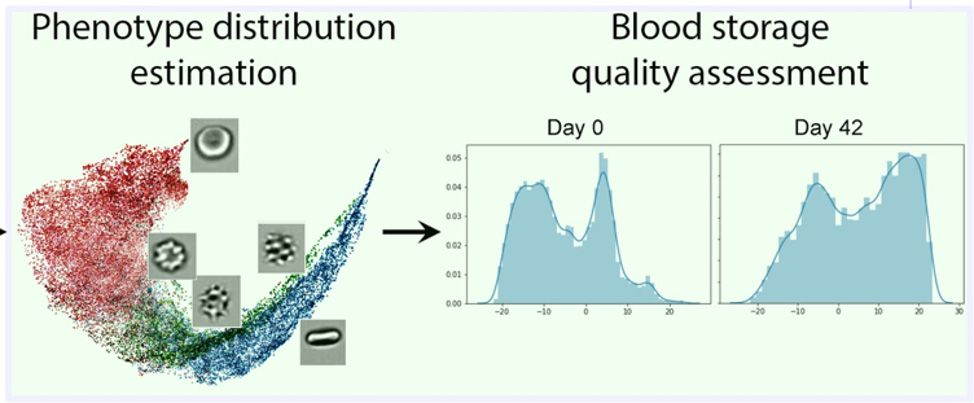
Figure 8. Distribution of phenotypes (morphologies) revealed during CNN visualization.
Plans for DeepFlow
Our group is currently evaluating several potential research projects that combine IFC and deep learning with MATLAB. One project builds upon the genotoxicity study but focuses on assessing the response of white blood cells in a patient who has undergone chemotherapy [3]. A second will extend DeepFlow to slide scan analysis, which could potentially enable companies to reanalyze large amounts of slide scan data. We are also developing a graphical interface for DeepFlow, which we will package with the network as a single, standalone application.
Published 2020
References
-
[1] Eulenberg, P., Köhler, N., Blasi, T. et al. “Reconstructing cell cycle and disease progression using deep learning.” Nature Communications 8, 463 (2017). doi: 10.1038/s41467-017-00623-3
-
[2] Doan, M., Sebastian, J.A. et. al. “Objective assessment of stored blood quality by deep learning.” Proceedings of the National Academy of Sciences Sep 2020, 117 (35) 21381-21390. doi: 10.1073/pnas.2001227117
-
[3] Doan, M., Case, M., Masic, D., Hennig, H., McQuin, C., Caicedo, J., Singh, S., Goodman, A., Wolkenhauer, O., Summers, H.D., Jamieson, D., van Delft, F.W., Filby, A., Carpenter, A.E., Rees, P. and Irving, J. (2020). “Label‐Free Leukemia Monitoring by Computer Vision.” Cytometry, 97: 407-414. doi: 10.1002/cyto.a.23987