Sequence Viewer
(Removed) Visualize and interactively explore biological sequences
The app has been removed.
Description
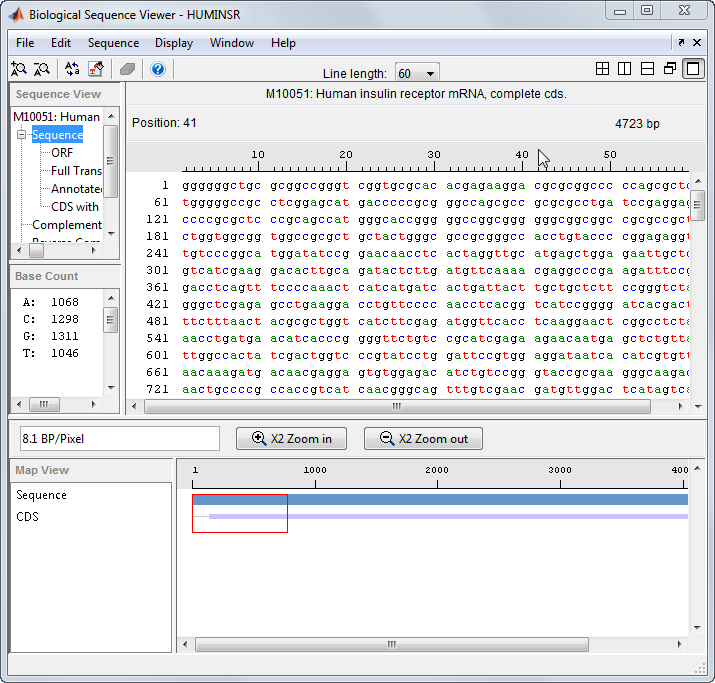
The Sequence Viewer app lets you visualize and explore amino acid or nucleotide sequences.
You can:
Import sequences from the NCBI or EMBL databases directly.
View and explore various sequence information (such as ORFs and CDSs) and basic statistics (such as percentages of base counts or amino acid counts).
View the complement and reverse complement sequences, and other sequence features such as genes and exons.
Extract protein coding sections of a nucleotide sequence and export them to the MATLAB® workspace.
Search for characteristic words or sequence patterns using regular expressions.
Open the Sequence Viewer App
MATLAB Toolstrip: On the Apps tab, under Computational Biology, click the app icon.
MATLAB command prompt: Enter
seqviewer.