getancestors
Find terms that are ancestors of specified Gene Ontology (GO) term
Syntax
Description
AncestorIDs = getancestors(GeneontObj,ID)GeneontObj, a geneont object, for GO terms
that are ancestors of the GO term(s) specified by ID, which is a GO term
identifier or vector of identifiers. The result AncestorIDs is a vector
of GO term identifiers including ID.
[
also returns the number of times each ancestor is found. AncestorIDs,Counts] = getancestors(GeneontObj,ID)Counts is a
column vector with the same number of elements as terms in
GeneontObj.
Tip
The Counts return value is useful when you tally counts in gene enrichment studies.
___ = getancestors(,
for any output arguments, specifies additional options using one or more name-value
arguments. For example, you can restrict the search to have up to two levels up in the gene
ontology by specifying GeneontObj,ID,Name,Value)AncestorIDs =
getancestors(GeneontObj,ID,Height=2).
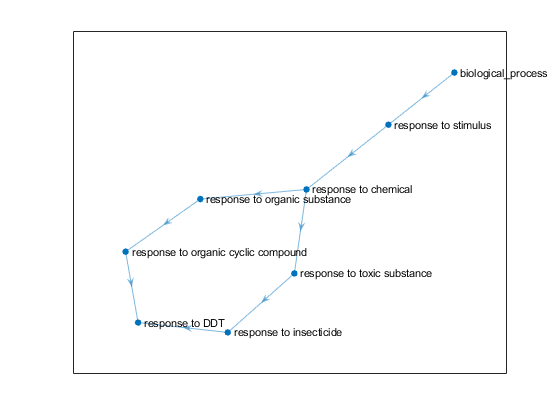
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced before R2006a
See Also
geneont | goannotread | num2goid | term