Explore a Protein Sequence Using the Sequence Viewer App
Overview of the Sequence Viewer
The Sequence Viewer app integrates many of the sequence functions in the Bioinformatics Toolbox™ toolbox. Instead of entering commands in the MATLAB® Command Window, you can select and enter options using the app.
Viewing Amino Acid Sequence Statistics
The following procedure illustrates how to view an amino acid sequence for an ORF located in a nucleotide sequence. You can import your own amino acid sequence, or you can get a protein sequence from the GenBank® database. This example uses the GenBank accession number NP_000511, which is the alpha subunit for a human enzyme associated with Tay-Sachs disease.
Select File > Download Sequence from > NCBI.
The Download Sequence from NCBI dialog box opens.
In the dialog box, type an accession number for an NCBI database entry, for example, NP_000511. Click the Protein option button, and then click OK.
The Sequence Viewer accesses the NCBI database on the Web and loads amino acid sequence information for the accession number you entered.
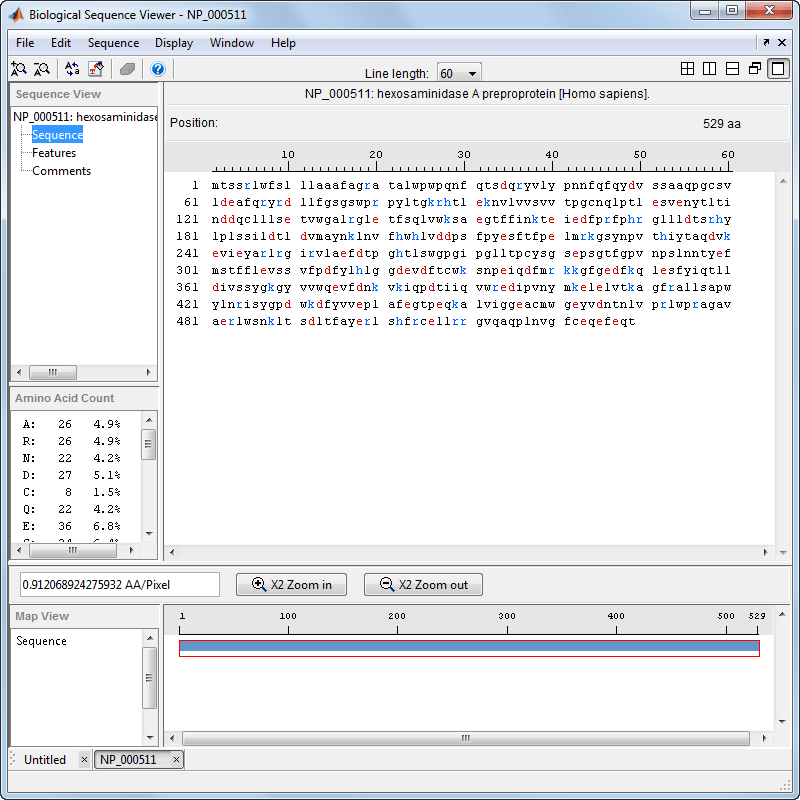
Select Display > Amino Acid Color Scheme, and then select Charge, Function, Hydrophobicity, Structure, or Taylor. For example, select Function.
The display colors change to highlight charge information about the amino acid residues. The following table shows color legends for the amino acid color schemes.
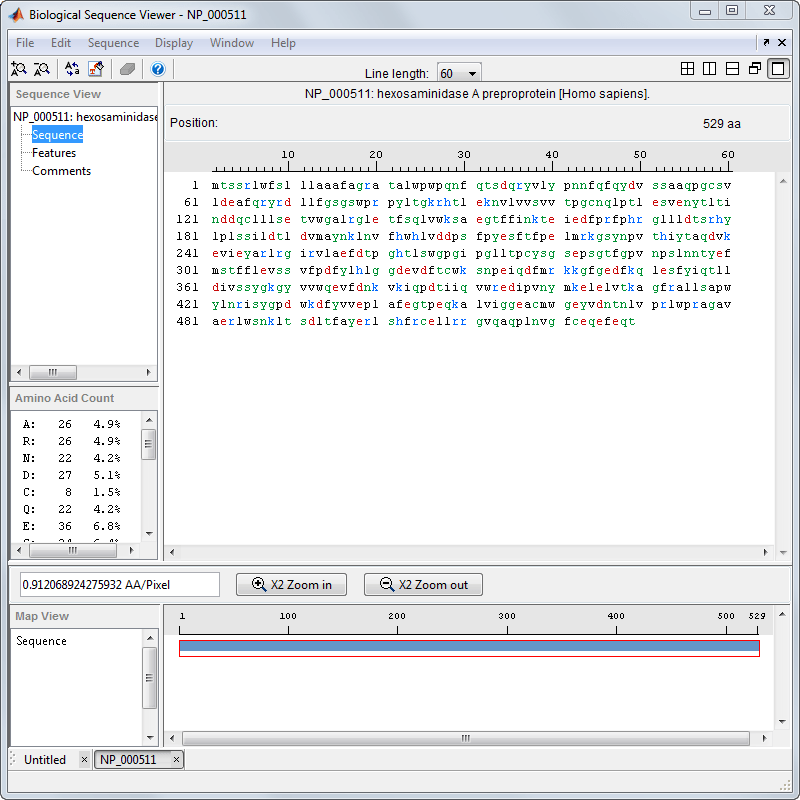
| Amino Acid Color Scheme | Color Legend |
|---|---|
| Charge |
|
| Function |
|
| Hydrophobicity |
|
| Structure |
|
| Taylor | Each amino acid is assigned its own color, based on the colors proposed by W.R. Taylor. |
Closing the Sequence Viewer
Close the Sequence Viewer from the MATLAB command line using the following syntax:
seqviewer('close')References
[1] Taylor, W.R. (1997). Residual colours: a proposal for aminochromography. Protein Engineering 10, 7, 743–746.