whiteningmat
Description
Examples
Apply the whiteningmat function to a 10-by-10 stationary colored noise covariance matrix. Plot the covariance matrix magnitude before and after whitening.
rng defaultSpecify the covariance matrix dimension N=10 and the number of independent noise samples M=1000.
N = 10; M = 1000;
Generate a Toeplitz matrix representing the covariance of stationary noise with a constant correlation structure.
corr_coeff = 0.6; alpha = 5; indices = 0:N-1; R = alpha*corr_coeff.^ abs(indices - indices');
Generate colored noise samples and the estimated original covariance matrix using Cholesky factorization.
L = chol(R,'lower');
x = L*randn(N,M);
R_est = cov(x');Plot estimated original covariance matrix.
figure
imagesc(R_est)
colorbar
title('Estimated Original Covariance Matrix Magnitude')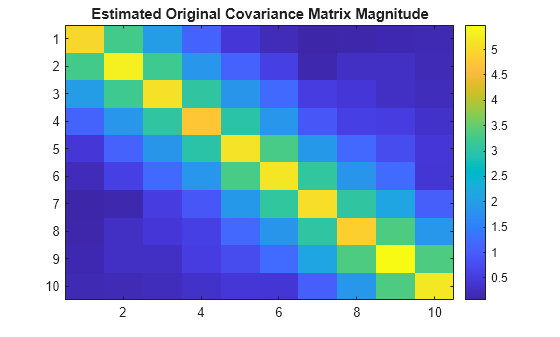
Apply the whitening transformation.
outnoisevar = 1; w = whiteningmat(R_est,OutputNoisePower=outnoisevar); x_whiten = w*x; R_whiten_est = cov(x_whiten.');
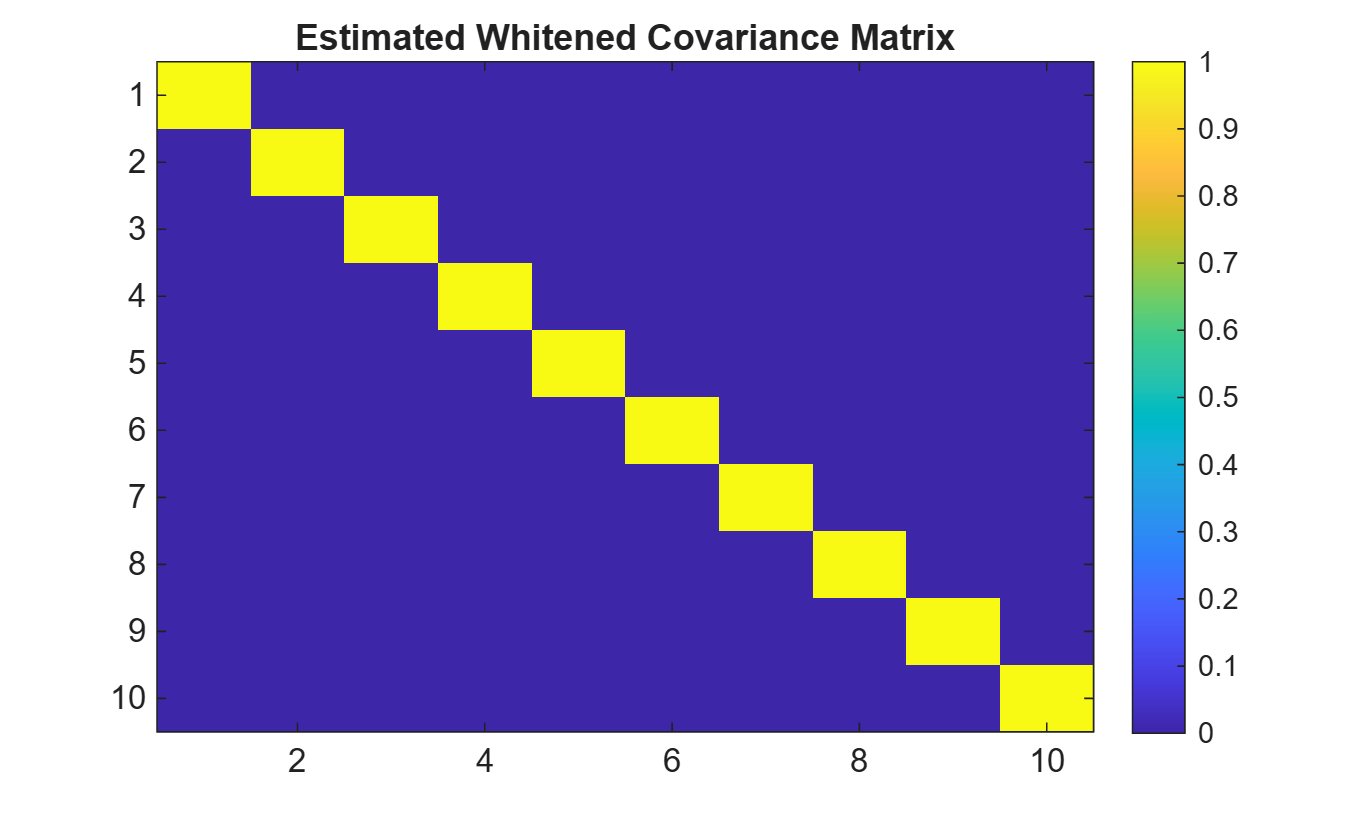
Plot the estimated whitened covariance matrix.
figure
imagesc(R_whiten_est)
colorbar
title('Estimated Whitened Covariance Matrix')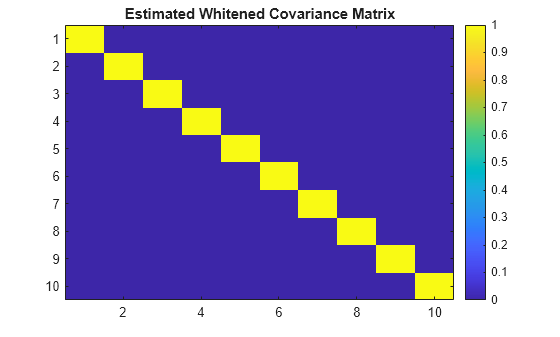
Apply the whiteningmat function to a 500-by-500 pink noise covariance matrix. Plot the covariance matrix magnitude before and after whitening.
rng defaultSpecify the covariance matrix dimension N = 500 and the number of independent channels M = 10000.
N = 500;
M = 1e4;
color = "pink";Configure a colored noise generator and then generate the noise.
pinkNoiseGen = dsp.ColoredNoise(Color=color, ...
SamplesPerFrame=N,NumChannels=M);
sig = pinkNoiseGen() + 1i*pinkNoiseGen();Plot estimated original covariance matrix.
covMatrix = cov(sig');
figure
imagesc(abs(covMatrix))
colorbar
title("Estimated Original Covariance Matrix Magnitude")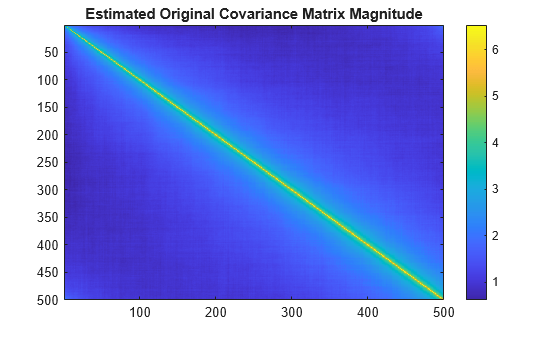
Apply the whitening transformation.
R = cov(sig'); W = whiteningmat(R); sigwhite = W*sig; covMatrixWhiten = cov(sigwhite');
Display the whitened covariance matrix.
figure
imagesc(abs(covMatrixWhiten))
colorbar;
title("Estimated Whitened Covariance Matrix Magnitude")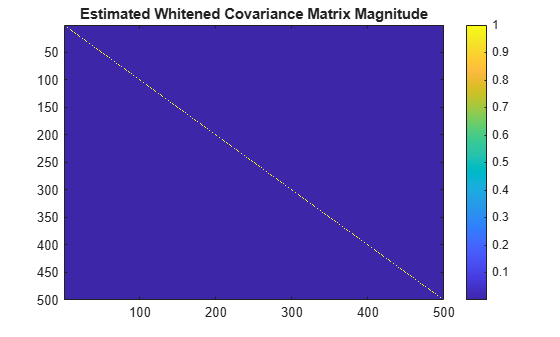
Generate M=1000 independent realizations of data contaminated by colored noise of dimension N=10. Apply whitening to the data before LRT detection to maintain a constant probability of false alarm. Display the probability of false alarm under the null hypothesis and the probability of detection under the alternative hypothesis.
rng default lrt = phased.LRTDetector(DataComplexity="Complex", ... ProbabilityFalseAlarm=0.1);
Create colored Gaussian noise.
N = 10;
M = 1000;
A = 10*rand(N);
R_noise = A'*A;
L_noise = chol(R_noise/2,"lower");Create the whitening matrix.
noisevar = 1; w = whiteningmat(R_noise,OutputNoisePower=noisevar);
Whiten the known reference signal.
xref = 5*ones(N,1); xref_whiten = w*xref;
Perform LRT detection under a null hypothesis.
x0 = L_noise*(randn(N,M) + 1i*randn(N,M)); x0_whiten = w*x0; dresult0 = lrt(x0_whiten,xref_whiten,noisevar);
Compute the probability of false alarm.
Pfa_whiten = sum(dresult0)/M; display(Pfa_whiten)
Pfa_whiten = 0.1030
Perform LRT detection under an alternative hypothesis
x1 = xref + L_noise*(randn(N,M) + 1i*randn(N,M)); x1_whiten = w*x1; dresult1 = lrt(x1_whiten,xref_whiten,noisevar);
Compute the probability of detection.
Pd_whiten = sum(dresult1)/M; display(Pd_whiten)
Pd_whiten = 0.6670
Consider two target signals impinging on a ULA of N=16 elements. The ULA also receives correlated clutter over different elements. Assume the covariance of clutter-plus-noise covariance matrix is known. Plot the MUSIC estimation result without applying whitening and plot the MUSIC estimation result after applying whitening.
rng default fc = 150.0e6; N = 16; array = phased.ULA(NumElements=N,ElementSpacing=1.0); estimator = phased.MUSICEstimator(SensorArray=array,... OperatingFrequency=fc,NumSignals=2, ... DOAOutputPort=true,NumSignalsSource="Property");
Generate the received signals.
fs = 8000;
t = (0:1/fs:1).';
sig1 = cos(2*pi*t*300.0);
sig2 = cos(2*pi*t*400.0);
az_true = [10, 60];
el_true = [20 -5];
sig = collectPlaneWave(array,[sig1 sig2], ...
[az_true; el_true],fc);Generate received clutter-plus-noise realizations.
corr_coeff = 0.9;
ula_indices = 0:N-1;
R = corr_coeff.^ abs(ula_indices - ula_indices');
U = chol(R/2,"upper");
clutter = (randn(size(sig)) + 1i*randn(size(sig)))*U;Perform MUSIC estimation before whitening.
[y,az_no_whiten] = estimator(sig + clutter); az_no_whiten = broadside2az(sort(az_no_whiten),[20 -5])
az_no_whiten = 1×2
0 9.0347
figure plotSpectrum(estimator,"NormalizeResponse",true) title("MUSIC Spectrum Without Whitening") tgtlabel = {"Target 1","Target 2"}; xline(az_true,"--",tgtlabel)
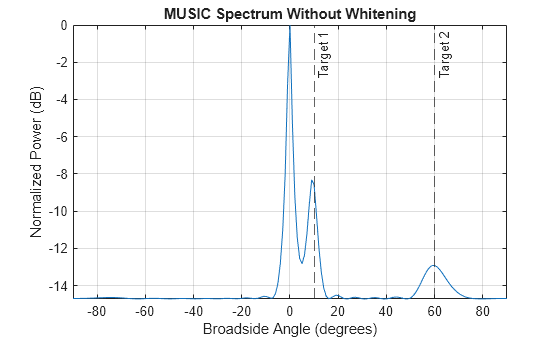
Perform MUSIC estimation after whitening.
w = whiteningmat(R); data_whiten = (sig + clutter)*w.'; [y_whiten,az_whiten] = estimator(data_whiten); az_whiten = broadside2az(sort(az_whiten),[20 -5])
az_whiten = 1×2
10.6490 60.3813
figure plotSpectrum(estimator,"NormalizeResponse",true) title("MUSIC Spectrum With Whitening") xline(az_true,"--",tgtlabel)
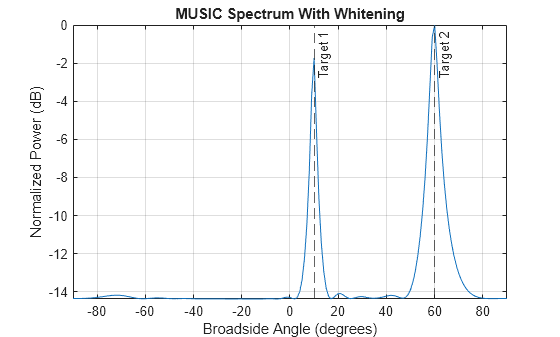
Consider two target signals impinging on a ULA of N=128 elements. The ULA also receives three interference signals. Assume the covariance of interference-plus-noise covariance matrix is known. Plot GLRT detection result without applying whitening and GLRT detection result with whitening applied before angle detection.
rng default glrt = phased.GLRTDetector(DataComplexity = "Complex", ... ProbabilityFalseAlarm = 0.01, ... ThresholdOutputPort = true);
Create two targets located at 11 and -47 degrees in azimuth.
tarAng = [11,-47]; tarNum = length(tarAng);
Create a ULA with 128 elements and half-wavelength element spacing.
N = 128;
fc = 3e8;
elementPos = (0:N-1)*physconst("LightSpeed")/fc/2;Create single-snapshot received signal from two targets at the ULA.
tarAmp = 1/sqrt(2)*(randn(tarNum,1)+1i*randn(tarNum,1)); signal = steervec(elementPos,tarAng)*tarAmp;
Create three interference sources located at -30, -19 and 50 degrees.
intAng = [-30,-19,50]; intNum = length(intAng);
Create single-snapshot complex colored Gaussian noise at the ULA.
npow = 0.1;
intAmp = 10/sqrt(2)*(randn(intNum,1)+1i*randn(intNum,1));
Aint= steervec(elementPos,intAng);
Rint = Aint*diag((abs(intAmp)).^2)*Aint';
Rint = (Rint+Rint')/2;
R = npow*eye(N) + Rint;
noise = chol(R/2,"lower")*(randn(N,1)+1i*randn(N,1));
x = signal + noise;Divide angle space into 256 angle bins.
D = 256; angGrid = asind(2*(-D/2:D/2-1)/D);
Perform GLRT detection without whitening.
obs = permute(steervec(elementPos,angGrid),[1 3 2]); hyp = [1,0]; [~,stat,th] = glrt(x,hyp,obs);
Perform GLRT detection with whitening.
w = whiteningmat(R); x_whiten = w*x; svs = w*steervec(elementPos,angGrid); obs_whiten = permute(svs,[1 3 2]); [~,stat_whiten,th_whiten] = glrt(x_whiten,hyp,obs_whiten);
Plot GLRT detection statistics at the 256 angle bins.
figure ax1 = subplot(2,1,1); plot(angGrid,pow2db(stat)) hold on grid on plot(angGrid,pow2db(th*ones(1,D))) title(ax1,"GLRT Detection Without Whitening") ax2 = subplot(2,1,2); plot(angGrid,pow2db(stat_whiten)) hold on grid on plot(angGrid,pow2db(th_whiten*ones(1,D))) title(ax2,"GLRT Detection With Whitening") ax = {ax1,ax2}; for idx = 1:2 xlabel(ax{idx},"Angle (degrees)") ylabel(ax{idx},"Power (dB)") tgtlabel = {"Target 1","Target 2"}; xline(ax{idx},tarAng,"--",tgtlabel) intlabel = {"Interference 1", ... "Interference 2","Interference 3"}; xline(ax{idx},intAng,"-.",intlabel) legend(ax{idx},{"Detection Statistics", ... "Detection Threshold"}) end
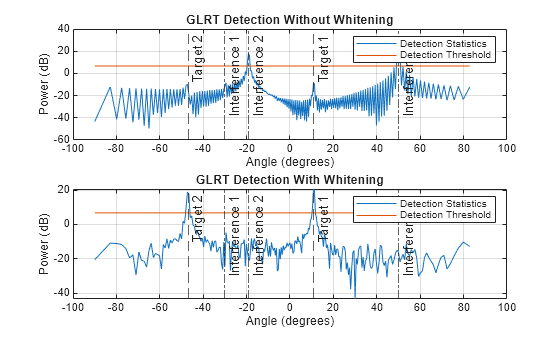
Input Arguments
Noise covariance, specified as an N-by-N real-valued positive definite matrix or N-by-N complex-valued positive definite matrix.
Data Types: single | double
Complex Number Support: Yes
Output power of whitened noise, specified as a positive real scalar.
Example: 10.0
Data Types: single | double
Output Arguments
Whitening transformation matrix, returned as an
N-by-N real-valued positive definite matrix or
an N-by-N complex-valued positive definite matrix.
The complexity of W matches the complexity of
ncov.
Data Types: single | double
Complex Number Support: Yes
Algorithms
This function uses Cholesky decomposition to whiten the covariance matrix
Ncov into a scaled identity matrix. See [1] or [2]. When applied to a
signal in noise it can be used to generate a transformed signal in white noise. Start with
the data model x = s + z, where x is a noise-free deterministic signal and
z is zero-mean Gaussian noise
specified by the covariance matrix Rn. You can input the noise covariance
matrix using the ncov argument. Applying a whitening matrix,
W to the data x, produces a transformed signal embedded in white noise.
The covariance of the whitened noise, represented as , is given by
where P is the white noise power specified by the
pow input argument. N represents the dimensions of
the noise matrix. When pow equals one, the whitened noise covariance
matrix is the identity matrix. For other positive values of pow the
whitened data covariance matrix is a scaled version of the identity matrix.
The images below show the magnitudes of the original unwhitened covariance matrix and
the whitened covariance matrix.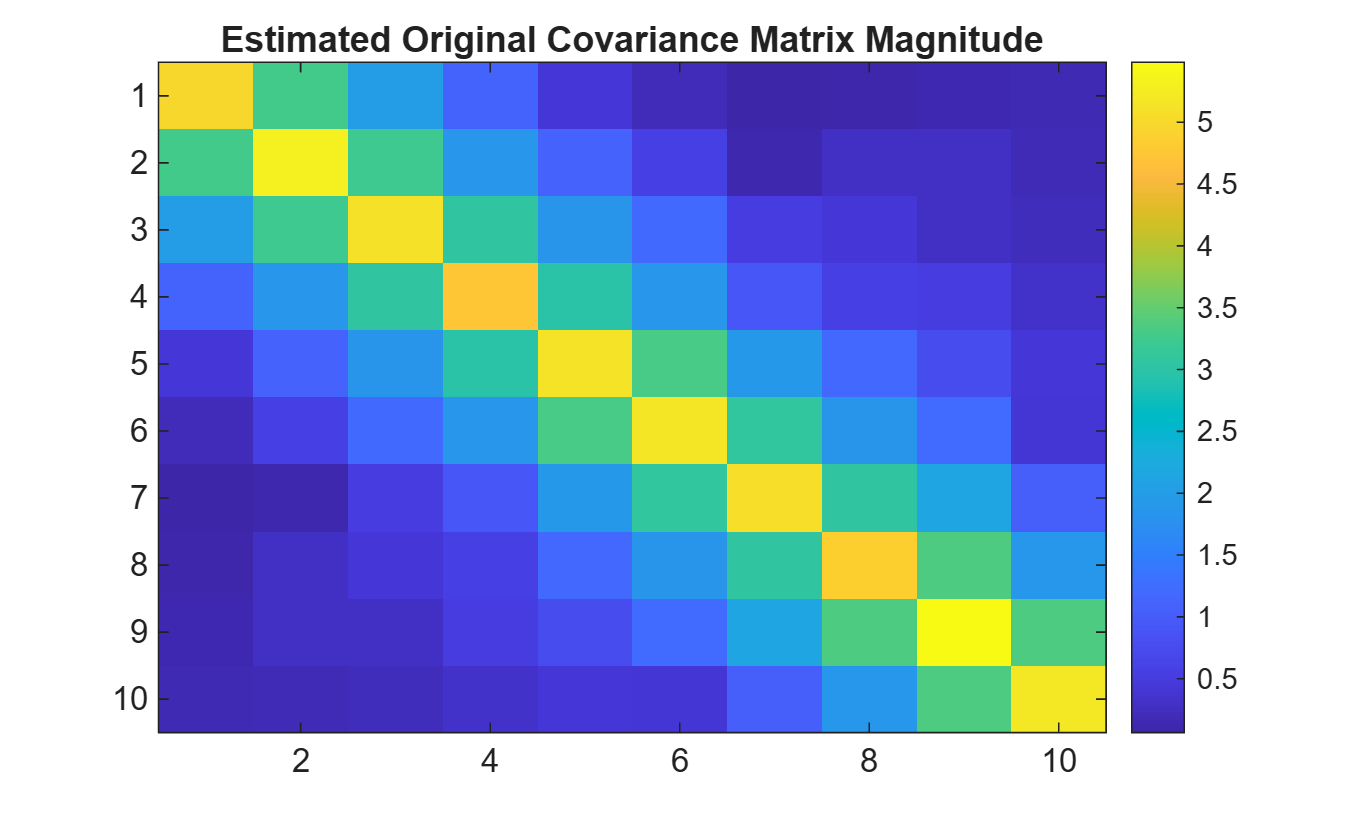
This function supports single and double precision for input arguments. If the input
argument ncov is single precision, the outputs are single precision. If
the input argument ncov is double precision, the outputs are double
precision. The precision of the outputs are independent of the precision of the other
arguments.
References
[1] Steven M. Kay, Fundamentals of Statistical Signal Processing, Detection Theory, 1998.
[2] Golub, Gene H. and Van Loan, Charles F. Matrix Computations - 4th Edition}, Philadelphia, PA: Johns Hopkins University Press, 2013.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)