加权非线性回归
此示例说明如何将非线性回归模型与具有非常量误差方差的数据进行拟合。
当测量误差的方差全部相同时,适合使用常规非线性最小二乘算法。当这个假设不成立时,适合使用加权拟合。此示例说明如何将权重与 fitnlm 函数一起使用。
要拟合的数据和模型
我们将使用收集的数据来研究工业和生活垃圾造成的水污染。这些数据在以下文献中有详细描述:Box, G.P., W.G. Hunter, and J.S. Hunter, Statistics for Experimenters (Wiley, 1978, pp. 483-487)。响应变量是以 mg/l 为单位的生化需氧量,预测变量是以天为单位的潜伏期。
x = [1 2 3 5 7 10]'; y = [109 149 149 191 213 224]'; plot(x,y,'ko') xlabel('Incubation (days), x') ylabel('Biochemical oxygen demand (mg/l), y')

我们假设已知前两个观测值的精度低于其余观测值。比如说,它们可能是用不同仪器测量的。对数据进行加权的另一个常见原因是记录的每个观测值实际上是在相同的 x 值处提取的几个测量值的均值。在此处的数据中,假设前两个值各代表一个原始测量值,其余四个值中的每个值分别代表一个从 5 个原始测量值求得的均值。那么,根据每次观察的测量次数进行加权就是合适的。
w = [1 1 5 5 5 5]';
不加权拟合模型
拟合这些数据的模型是一条缩放的指数曲线,当 x 变大时,曲线变为水平。
modelFun = @(b,x) b(1).*(1-exp(-b(2).*x));
仅根据粗略的视觉拟合就可以看出,在 x = 15 附近,值大约为 240 的位置,沿这些点绘制的曲线似乎就会变平。因此,我们使用 240 作为 b1 的起始值,而且由于 e^(-.5*15) 比 1 小,所以我们使用 .5 作为 b2 的起始值。
start = [240; .5];
忽略测量误差的危险在于拟合可能受不精确测量值的过度影响,因而可能无法很好地拟合已知精确的测量值。我们先进行不加权拟合,然后将数据与各点进行比较。
nlm = fitnlm(x,y,modelFun,start); xx = linspace(0,12)'; line(xx,predict(nlm,xx),'linestyle','--','color','k')

请注意,拟合曲线靠近前两个点,但似乎偏离了其他点的趋势。
加权拟合模型
现在我们再尝试进行加权拟合。
wnlm = fitnlm(x,y,modelFun,start,'Weight',w)wnlm =
Nonlinear regression model:
y ~ b1*(1 - exp( - b2*x))
Estimated Coefficients:
Estimate SE tStat pValue
________ ________ ______ __________
b1 225.17 10.7 21.045 3.0134e-05
b2 0.40078 0.064296 6.2333 0.0033745
Number of observations: 6, Error degrees of freedom: 4
Root Mean Squared Error: 24
R-Squared: 0.908, Adjusted R-Squared 0.885
F-statistic vs. zero model: 696, p-value = 8.2e-06
line(xx,predict(wnlm,xx),'color','b')
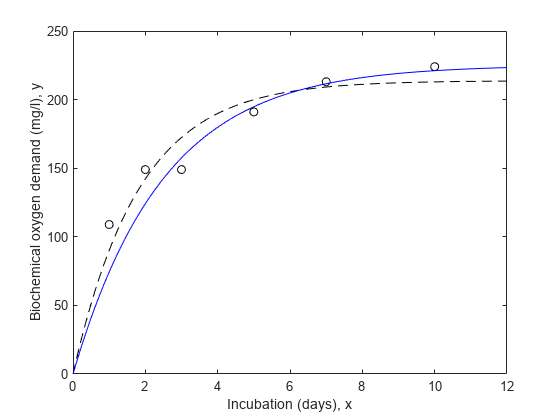
在这种情况下,估计的总体标准差描述了权重或测量精度为 1 的“标准”观测值的平均变化。
wnlm.RMSE
ans = 24.0096
对于任何分析来说,估计模型拟合的精度都是重要的一部分。系数显示了参数的标准误差,但我们也可以计算参数的置信区间。
coefCI(wnlm)
ans = 2×2
195.4650 254.8788
0.2223 0.5793
估计响应曲线
接下来,我们将计算参数的拟合响应值和置信区间。默认情况下,这些宽度表示预测值的逐点置信边界,但我们将请求整个曲线的同时置信区间。
[ypred,ypredci] = predict(wnlm,xx,'Simultaneous',true); plot(x,y,'ko',xx,ypred,'b-',xx,ypredci,'r:') xlabel('x') ylabel('y') ylim([-150 350]) legend({'Data','Weighted fit','95% Confidence Limits'}, ... 'location','SouthEast')
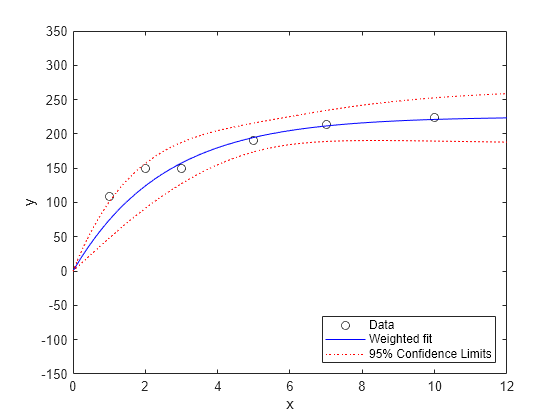
请注意,两个下加权点并没有像其他点一样很好地与曲线拟合。这符合您对加权拟合的预期。
您还可以估计以后在指定 x 值处所得观测值的预测区间。计算这些区间时,会假定权重或测量精度为 1。
[ypred,ypredci] = predict(wnlm,xx,'Simultaneous',true, ... 'Prediction','observation'); plot(x,y,'ko',xx,ypred,'b-',xx,ypredci,'r:') xlabel('x') ylabel('y') ylim([-150 350]) legend({'Data','Weighted fit','95% Prediction Limits'}, ... 'location','SouthEast')

权重的绝对尺度实际上不影响参数估计值。按任何常量重新调整权重会得到相同的估计值。但它们确实影响置信边界,因为边界代表权重为 1 的观测值。这里您可以看到,与置信界限相比,权重较大的点似乎太靠近拟合线。
假设我们对一个新的观测值感兴趣,它基于五个测量值的均值,就像此图中后四个点一样。通过使用 predict 函数的 Weights 名称-值参量指定观测值权重。
[new_ypred,new_ypredci] = predict(wnlm,xx,'Simultaneous',true, ... 'Prediction','observation','Weights',5*ones(size(xx))); plot(x,y,'ko',xx,new_ypred,'b-',xx,new_ypredci,'r:') xlabel('x') ylabel('y') ylim([-150 350]) legend({'Data','Weighted fit','95% Prediction Limits'}, ... 'location','SouthEast')

predict 函数通过 MSE*(1/W(i)) 估计观测值 i 处的误差方差,其中 MSE 是均方误差。因此,置信区间变窄。
残差分析
除了绘制数据和拟合之外,还会绘制拟合残差对预测变量的图,以诊断模型的任何问题。残差应该表现为独立同分布 (i.i.d.),但方差与权重的倒数成正比。绘制标准化残差图,以确认这些值是具有相同方差的 i.i.d.。标准化残差是原始残差除以其估计标准差。
r = wnlm.Residuals.Standardized; plot(x,r,'b^') xlabel('x') ylabel('Standardized Residuals')

可以看出,此残差图存在一定的系统模式。请注意最后四个残差的线性趋势,这表明模型可能不会随着 x 增加而快速增加。而且,残差的模随 x 增大而减小,这表明测量误差可能取决于 x。这些特性值得进一步研究,但由于数据点太少,不足以确定这些明显模式的研究价值。
另请参阅
NonLinearModel | fitnlm | predict | coefCI