aacount
Count amino acids in sequence
Description
countStruct = aacount(SeqAA)SeqAA, an amino acid
sequence, and returns the counts in countStruct, a 1-by-1 MATLAB® structure containing fields for the standard 20 amino acids
(A, R, N, D,
C, Q, E, G,
H, I, L, K,
M, F, P, S,
T, W, Y, and
V).
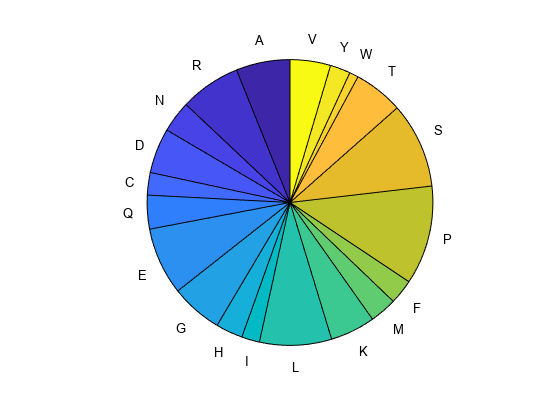
countStruct = aacount(SeqAA,Name=Value)countStruct = aacount(SeqAA,Chart="pie") creates a pie chart showing
relative proportions of the amino acids.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced before R2006a
See Also
aminolookup | atomiccomp | basecount | codoncount | dimercount | isoelectric | molweight | proteinplot | proteinpropplot