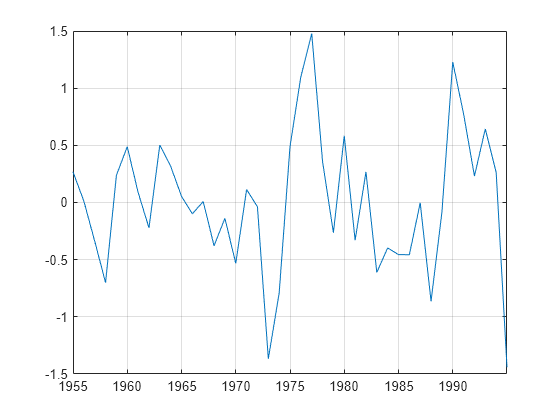egcitest
Engle-Granger cointegration test
Syntax
Description
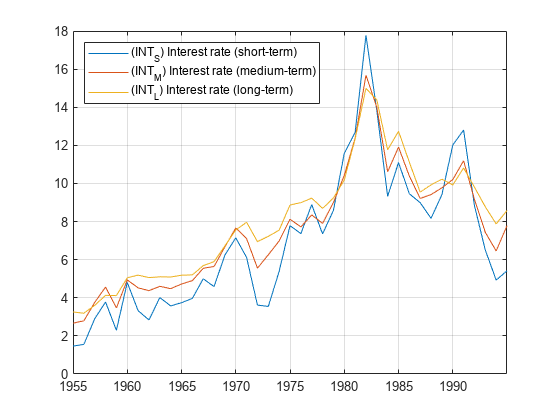
h = egcitest(Y)egcitest forms test statistics by regressing the
response data Y(:,1) onto the predictor data
Y(:,2:end), and then it tests the residuals for a unit root.
StatTbl = egcitest(Tbl)
The response variable in the regression is the first table variable, and all other
variables are the predictor variables. To select a different response variable for the
regression, use the ResponseVariable name-value argument. To select
different predictor variables, use the PredictorNames name-value
argument.
[___] = egcitest(___,
specifies options using one or more name-value arguments in
addition to any of the input argument combinations in previous syntaxes.
Name=Value)egcitest returns the output argument combination for the
corresponding input arguments.
Some options control the number of tests to conduct. The following conditions apply when
egcitest conducts multiple tests:
For example, egcitest(Tbl,ResponseVariable="GDP",Alpha=0.025,Lags=[0
1]) chooses GDP as the response variable from the table
Tbl and conducts two tests at a level of significance of 0.025. The
first test includes 0 lag in the residual regression, and the second test
includes 1 lag in the residual regression.
[___,
additionally returns the following structures of regression statistics, which are required
to form the test statistic:reg1,reg2] = egcitest(___)
reg1– Statistics resulting from the cointegrating regression of the specified response variableResponseVariableonto the specified predictor variablesPredictorVariablesreg2– Statistics resulting from the residual regression implemented by the specified unit root testRReg
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Tips
To draw valid inferences from the test, determine a suitable value for
Lags. For more details, see theadftestTips and thepptestTips.Samples with less than approximately 20 through 40 observations (depending on the dimension of the data
numDims) can yield unreliable critical values, and therefore unreliable inferences. See [3].If a test result suggests that the time series are cointegrated, you can use the residuals as data for the error-correction term in a VEC representation of the variables. Follow this procedure:
Alternative Functionality
App
The Econometric Modeler app enables you to conduct the Engle-Granger cointegration test.
References
[2] Hamilton, James D. Time Series Analysis. Princeton, NJ: Princeton University Press, 1994.
Version History
Introduced in R2011a