corr
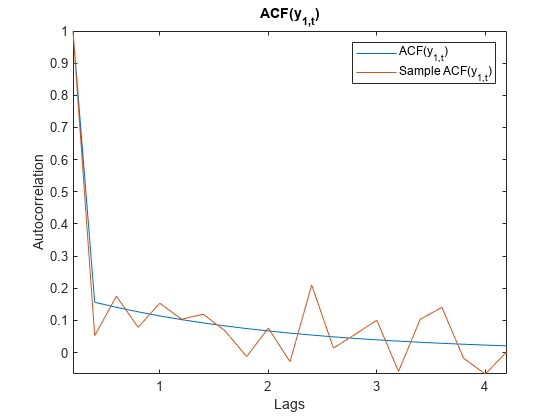
Model-implied temporal correlations of state-space model
Syntax
Description
The corr function returns model-implied temporal correlations and covariances of the state or measurement variables in a stationary, time-invariant state-space model. To determine whether the model captures characteristics present in the data, You can compare model-implied associations of present and lagged variables to sample analogues. Other state-space model tools to characterize the dynamics of a specified system include the following:
The impulse response function (IRF), computed by
irfand plotted byirfplot, traces the effects of a shock to a state disturbance on the measurement variables in the system.The forecast error variance decomposition (FEVD), computed by
fevd, provides information about the relative importance of each state disturbance in affecting the forecast error variance of all measurement variables in the system.
Fully Specified State-Space Model
Cyy = corr(Mdl,Name,Value)'Covariance',true,'NumLags',10 specifies returning temporal covariances Cov(yt,yt – h), h = 0 through 10.
[ also returns Corr(xt,xt – h), the correlations between the state variables and their self-lags Cyy,Cxx,Cyx] = corr(___)Cxx, and Corr(yt,xt – h), the correlations between the state variables and their self lags Cxx and the cross-correlations between the measurement variables and lags of the state variables Cyx using any of the input argument combinations in the previous syntaxes. h is the value of the NumLags name-value argument. corr returns covariances when the value of the Covariance name-value argument is true.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Tips
To obtain an association matrix of lead variables from an association matrix of lagged variables, use the identity
where:
C is an association function, either Corr or Cov.
at and bt are yt or xt.
Version History
Introduced in R2021a