Optimize Simulink Model in Parallel with Optimization Explorer
This example shows how to optimize a Simulink® model in parallel using the Optimization Explorer app. To optimize a Simulink model using Optimization Explorer, you typically must use serial computing, not parallel computing. This limitation is due to the design of optimization solvers, which generally compute in parallel by using the parfor or parfeval function. Simulink models do not support these functions for parallel computation.
However, as shown in the example Optimize Simulink Model in Parallel, you can use Global Optimization Toolbox solvers that support vectorized function evaluation to optimize a Simulink model in parallel. These solvers are ga, patternsearch, surrogateopt, and particleswarm.
To set up Optimization Explorer for parallel optimization of a Simulink model, you start by writing a vectorized objective function based on the model. Then you specify attributes for the vectorized objective function in Optimization Explorer, and choose solvers and options.
Vectorized Objective Function Based on Simulink Model
To optimize a Simulink model in parallel, write a vectorized objective function that:
Accepts a matrix as input, where each row of the matrix is a vector of parameters
Performs simulations in parallel by calling the
parsim(Simulink) functionReturns a vector of values, where each entry in the vector contains the value of the objective function corresponding to the parameters in one row of the input matrix
This figure shows how a vectorized objective function works with a matrix of input values, giving a column vector of output values.

For this example, use the Lorenz_system.slx Simulink model.
The objective function is to minimize the sum of squares of the difference between the Lorenz system and the uniform circular motion over a set of times from 0 through 1/10. For times xdata, the objective function is
objective = sum((fitlorenzfn(x,xdata) - lorenz(xdata)).^2) - (F(1) + F(end))/2
where:
The
fitlorenzfnhelper function (shown at the end of this example) computes points on the uniform circular motion path.lorenz(xdata)represents the 3-D evolution of the Lorenz system at timesxdata.Frepresents the vector of squared distances between corresponding points in the circular and Lorenz systems.The objective subtracts half of the values at the endpoints to best approximate an integral.
Enter the data for the uniform circular motion.
x = zeros(8,1); x(1) = 1.2814; x(2) = -1.4930; x(3) = 24.9763; x(4) = 14.1870; x(5) = 0.0545; x(6:8) = [13.8061;1.5475;25.3616];
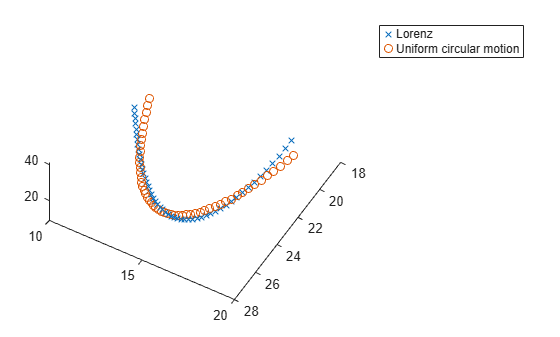
Load the Lorenz system and simulate it with the base parameters. Create a 3-D plot of the resulting system and compare the plot with the specified uniform circular motion.
model = "Lorenz_system"; open_system(model); in = Simulink.SimulationInput(model); % params [X0,Y0,Z0,Sigma,Beta,Rho] params = [10,20,10,10,8/3,28]; % Original parameters Sigma, Beta, Rho in = setparams(in,model,params); out = sim(in); yout = out.yout; tlist = yout{1}.Values.Time; YDATA = fitlorenzfn(x,tlist); figure; plot3(yout{1}.Values.Data,yout{2}.Values.Data,yout{3}.Values.Data,"x"); view([-30 -70]) hold on plot3(YDATA(:,1),YDATA(:,2),YDATA(:,3),"o") legend("Lorenz","Uniform circular motion") hold off
The parobjfun helper function shown below accepts a matrix of parameters, where each row of the matrix represents one set of parameters. The function calls the setparams helper function (shown at the end of this example) to set parameters for a Simulink.SimulationInput vector. The parobjfun function then calls parsim to evaluate the model on the parameters in parallel. The parobjfun function:
Accepts a vector of parameters with N elements
Creates a vector of N simulation inputs for the model
Initializes the output as a vector of N zeros
Maps the optimization parameters to each simulation input
Sets the
parsimproperties and performs the simulation in parallelFetches the simulation outputs (a blocking call) and computes the final outputs
function f = parobjfun(params,YDATA,model) N = size(params,1); simIn(1:N) = Simulink.SimulationInput(model); f = zeros(N,1); for i = 1:N simIn(i) = setparams(simIn(i),model,params(i,:)); end simOut = parsim(simIn,ShowProgress="off", ... UseFastRestart="on"); for i = 1:N yout = simOut(i).yout; vals = [yout{1}.Values.Data,yout{2}.Values.Data,yout{3}.Values.Data]; g = sum((YDATA - vals).^2,2); f(i) = sum(g) - (g(1) + g(end))/2; end end
Set bounds for solvers 20% above and below the current parameter values.
lb = 0.8*params; ub = 1.2*params;
For added speed, set the simulation to use fast restart.
set_param(model,FastRestart="on");Define a function to close the model using the closeModel helper function (shown at the end of this example).
c = onCleanup(@()closeModel(model));
For reproducibility, set the random stream.
rng(1)
Define the objective function as parobjfun with its extra parameters.
fitobjective = @(params)parobjfun(params,YDATA,model);
Vectorized Objective Function in Optimization Explorer
Start Optimization Explorer by entering the following at the command line:
optimizationExplorer
Because you need to use vectorized function evaluations, specify the problem using Solver Inputs.
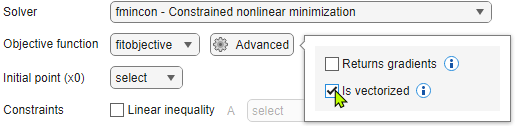
For the Objective function, select fitobjective. Click Advanced and select Is vectorized. You can leave the Solver selection at the default setting; you specify solvers in the next section.
For this problem, choose params as the Initial point (x0) value. For the Constraints, select Bounds and set the bounds lb and ub to lb and ub, respectively. Click Next to continue.
Manually Schedule Solver Configurations
To optimize a Simulink model in parallel, you must run manual configurations, not auto configurations. Therefore, you do not need to select any of the problem attributes such as Convex, Nonsmooth, Expensive, or Simulation. Click Next to continue.
Specify to run Optimization Explorer in Manual mode.

Click Start Session.
Leave Parallel Off
Although you will run Simulink in parallel in Optimization Explorer, do not use the app's default parallel mode. The Use Parallel button is not appropriate for optimizing a Simulink model.

Choose Solvers and Options
To schedule solver configurations, you use the Select Solvers section of the toolstrip. After you select a solver, the app opens a tab with options for that solver. Begin by selecting the ga solver.
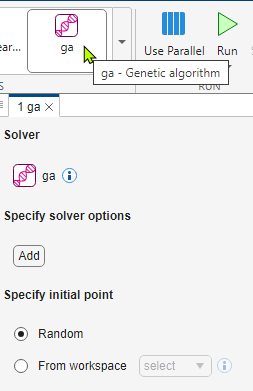
Use the Add button to display solver options you can set. For some solvers, such as patternsearch, you first need to select the algorithm for the solver. To set additional options, click the plus sign that appears to the right of an option you set.
Schedule these solver algorithms, all of which can run in a vectorized manner: ga, patternsearch (nups), patternsearch (classic), surrogateopt, and particleswarm. To run the example reasonably quickly, set the maximum number of function evaluations to 300 for each solver (for ga and particleswarm, have a population of 12 run for 25 iterations). Set the following options:
ga
Population settings > Population size
12Run time limits > Max generations
25Specify initial point: From workspace >
params
patternsearch
Algorithm >
nupsRun time limits > Max function evals
300Specify initial point: From workspace >
params
patternsearch
Algorithm >
classicRun time limits > Max function evals
300Poll settings > Do a complete poll Select the check box
Note: To run patternsearch (classic) in a vectorized manner, you must use a complete poll.
4. Specify initial point: From workspace > params
surrogateopt
Algorithm settings > Batch size for surrogate update and function evaluation
12Run time limits > Max function evals
300Specify initial point: From workspace >
params
particleswarm
Run time limits > Max iterations
25Swarm settings > Swarm size
12Specify initial point: From workspace >
params
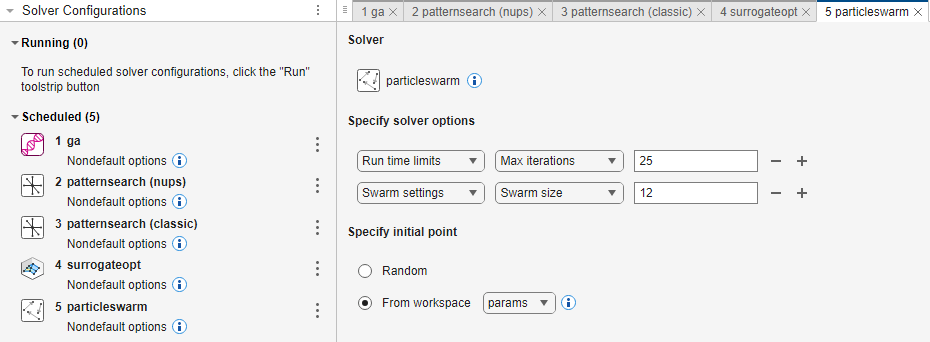
After making the solver selections, run Optimization Explorer by clicking the Run button on the toolstrip. The Solver Configurations panel shows which solver is running and which solvers are scheduled to run.
Results
After Optimization Explorer finishes, you see the following or similar results.
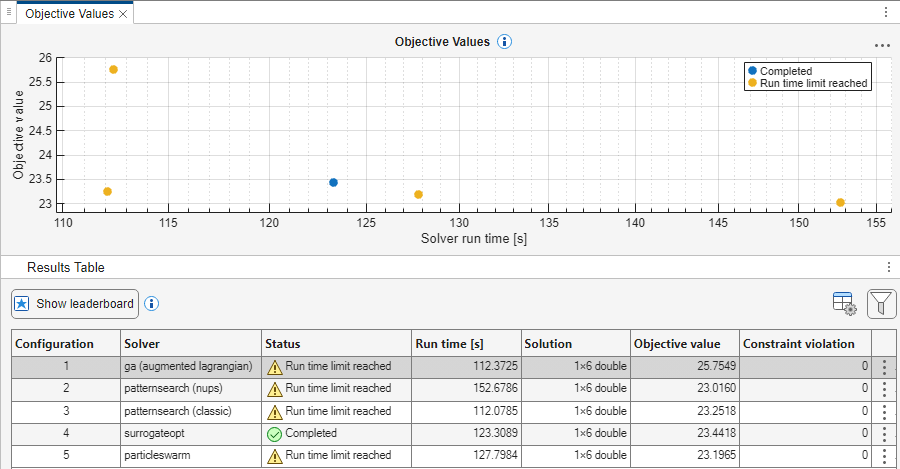
As shown in these results:
The best (smallest) objective function value is about
23.016, and is returned by thepatternsearch(nups)solver.The fastest run times are about
112seconds for both thegasolver and thepatternsearch(classic)solver.All solvers except
gareach an objective function value of less than 24. All solvers start from the objective function valuefitobjective(params)=26.9975.
Helper Functions
This code creates the fitlorenzfn helper function.
function f = fitlorenzfn(x,xdata) % Lorenz function theta = x(1:2); R = x(3); V = x(4); t0 = x(5); delta = x(6:8); f = zeros(length(xdata),3); f(:,3) = R*sin(theta(1))*sin(V*(xdata - t0)) + delta(3); f(:,1) = R*cos(V*(xdata - t0))*cos(theta(2)) ... - R*sin(V*(xdata - t0))*cos(theta(1))*sin(theta(2)) + delta(1); f(:,2) = R*sin(V*(xdata - t0))*cos(theta(1))*cos(theta(2)) ... - R*cos(V*(xdata - t0))*sin(theta(2)) + delta(2); end
This code creates the setparams helper function.
function pp = setparams(in,model,params) % parameters [X0,Y0,Z0,Sigma,Beta,Rho] pp = in.setVariable("X0",params(1),"Workspace",model); pp = pp.setVariable("Y0",params(2),"Workspace",model); pp = pp.setVariable("Z0",params(3),"Workspace",model); pp = pp.setVariable("Sigma",params(4),"Workspace",model); pp = pp.setVariable("Beta",params(5),"Workspace",model); pp = pp.setVariable("Rho",params(6),"Workspace",model); end
This code creates the closeModel helper function. To close the model, you must first disable fast restart.
function closeModel(model) set_param(model,FastRestart="off"); close_system(model) end