Medical Image Labeler
Interactively explore, label, and publish animations of 2-D or 3-D medical image data
Since R2022b
Description
The Medical Image Labeler app enables you to label ground truth data in medical images. Using the app, you can:
Import multiple 2-D images or 3-D image volumes.
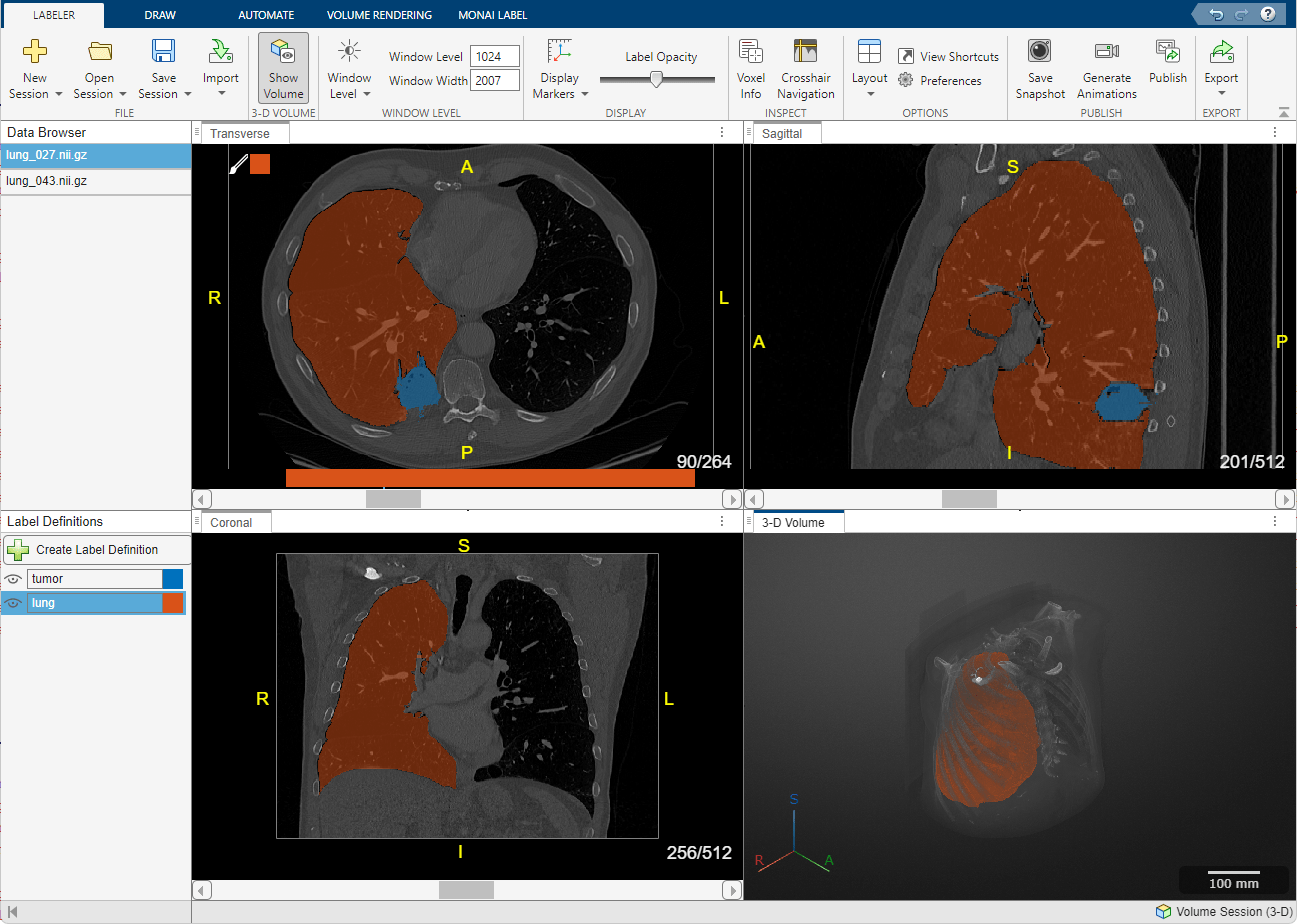
Explore images as slice planes or volumes with anatomical orientation markers and scale bars. Navigate volume slices using crosshairs.
Publish 2-D and 3-D PNG or PDF image snapshots and GIF animations.
Create multiple pixel label definitions to label regions of interest. Label pixels using automatic algorithms such as flood fill, semi-automatic techniques such as interpolation, and manual techniques such as painting by superpixels.
Segment and label objects in medical images and cross-sections of medical volumes using the Medical Segment Anything Model (MedSAM) algorithm. This functionality requires the Medical Imaging Toolbox™ Model for Medical Segment Anything Model support package.
Connect to the Medical Open Network for AI (MONAI) Label platform to apply fully automated and interactive deep learning models for segmenting radiology images. This functionality requires the Medical Imaging Toolbox Interface for MONAI Label Library support package.
Write, import, and use your own custom automation algorithm to automatically label ground truth data.
Export the labeled ground truth data as a
groundTruthMedicalobject. You can use this object to share labels with colleagues or for training semantic segmentation deep learning networks.
The Medical Image Labeler app supports 2-D images and image sequences stored in
the DICOM format, as well as the formats in the imformats registry. An image sequence is a series of images related by time,
such as ultrasound data. You can also import 2-D images or image series from
medicalImage objects in the workspace. The app supports 3-D image volume
data stored in the DICOM (single or multifile volume), NIfTI, NRRD, and TIFF file formats. You
can also import 3-D image volumes from medicalVolume objects in the
workspace.
To learn more about this app, see Get Started with Medical Image Labeler.
Note
The app is not supported in MATLAB® Online™. For details, see Specifications and Limitations.
Open the Medical Image Labeler App
MATLAB Toolstrip: On the Apps tab, under Image Processing and Computer Vision, click the Medical Image Labeler app icon.
MATLAB command prompt: Enter
medicalImageLabeler.
Examples
- Get Started with Medical Image Labeler
- Visualize 3-D Medical Image Data Using Medical Image Labeler
- Label 2-D Ultrasound Series Using Medical Image Labeler
- Label 3-D Medical Image Using Medical Image Labeler
- Automate Labeling in Medical Image Labeler
- Get Started with MedSAM in Medical Image Labeler
- Get Started with MONAI Label in Medical Image Labeler
- Segment CT Scan Using MONAI Label
- Collaborate on Multi-Labeler Medical Image Labeling Projects
Programmatic Use
Tips
When you import a volume from a DICOM file whose metadata specifies the
ResaleInterceptandRescaleSlopemetadata attributes, the app imports values in Hounsfield Units. This behavior is consistent with how themedicalVolumereads DICOM values in theVoxelsproperty.
Version History
Introduced in R2022bSee Also
Apps
- Medical Volume Viewer | Medical Registration Estimator | Image Labeler (Computer Vision Toolbox)