Tumor grading in GUI or ...?
5 次查看(过去 30 天)
显示 更早的评论
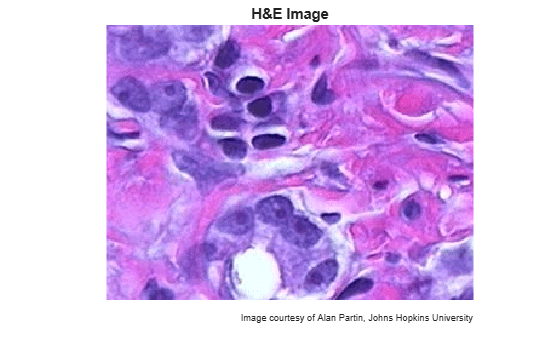
I have micro tumor images(9) and there are areas of dlbcl tumor (Nd DAB-stained and Nh H-stained) Proliferation index for this is a solution of the equation PIc=Nd/(Nh+Nd). I have to find PIc for all pictures knowing RGB intervals for Nh= (<45, 180>, <50, 185>, <160, 215>) and for Nd= (<40, 115>, <6,80>, <10,75>). How AM i supposed to do it? Please help!!
4 个评论
Image Analyst
2018-11-22
And the equations in my answer below don't do it? Please explain why not. And attach your image.
回答(4 个)
Image Analyst
2018-11-22
Use the code in Jan's comment above, but modify it to get two binary images: matchNh and matchNd. Then I think you want the sum (count) and ratio
Nd = nnz(matchNd)
Nh = nnz(matchNh)
Plc = Nd / (Nh + Nd)
0 个评论
Image Analyst
2018-11-22
4 个评论
Image Analyst
2018-11-23
Look at the gamuts - they're essentially a 3-D histogram where there is one point for every (r,g,b) triplet (every pixel value/color).
Point 1: note how there are no well contained clusters - it's basically a continuum - so the dividing plane will essentially be sort of arbitrary. You could almost put it wherever you want but at least kmeans has some sort of logic behind where it splits the gamut into the 2 colors.
Point 2: thresholding essentially divides up the gamut by slicing the color could with planes aligned with the axes, and you can see that there are no planes perpendicular to the R, G, and B axes that will do even a half way decent job of dividing the image up into different colors that are meaningfull. With HSV color space, your dividing surfaces are basically cones, sectors, and planes perpendicular to the V axis. By doing that, you can definitely carve out different regions because the hue, saturation, and brightness are different between the pink, purple, and white. The R, G, and B are also different, but you can see that planes perpendicular to the axes will not carve that cloud up into pink, purple, and white sub-clouds.
Aneta Chwala
2018-11-27
3 个评论
Image Analyst
2018-11-27
Yes, you may.
If it's unrelated to your first one way at the top, then create a new thread.
另请参阅
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!