loss
Description
L = loss(mdl,tbl,ResponseVarName)L for the compact classification XGBoost model ens
using the predictor data in table tbl and the true class labels in
tbl.ResponseVarName. The interpretation of L
depends on the loss function (LossFun) and weighting scheme
(Weights). In general, better classifiers yield smaller
classification loss values. The default LossFun value is
"classiferror" (misclassification rate in decimal).
L = loss(___,Name=Value)
Examples
Import a pretrained XGBoost classification model trained using the ionosphere dataset. The pretrained model is provided with this example.
load ionosphere modelfile = "trainedXGBoostModel.json"; Mdl = importModelFromXGBoost(modelfile)
Mdl =
CompactClassificationXGBoost
ResponseName: 'Y'
ClassNames: [0 1]
ScoreTransform: 'logit'
NumTrained: 30
ImportedModelParameters: [1×1 struct]
Properties, Methods
The model is imported as a CompactClassificationXGBoost model object.
Convert the response data to a boolean array to match the imported model.
Y = (Y=="g");Use the loss function to evaluate the classification cost of the model.
loss(Mdl,X,Y,LossFun="classifcost")ans = single
0.0476
Input Arguments
Compact classification XGBoost model, specified as a CompactClassificationXGBoost model object created with importModelFromXGBoost.
Sample data, specified as a table. Each row of tbl corresponds to one observation, and each column corresponds to one predictor variable. tbl must contain all of the predictors used to train the model. Multicolumn variables and cell arrays other than cell arrays of character vectors are not allowed.
Data Types: table
Response variable name, specified as the name of a variable in tbl. If mdl.ResponseName is the response variable name, then you do not need to specify ResponseVarName.
If you specify ResponseVarName, you must specify it as a character vector or string scalar. For example, if the response variable Y is stored as tbl.Y, then specify it as "Y". Otherwise, the software treats all columns of tbl, including Y, as predictors.
The response variable must be a logical or numeric vector.
Data Types: char | string
Class labels, specified as a logical or numeric vector. Y must have the same data type as tbl or X.
Y must be of the same type as the classification used to train mdl, and its number of elements must equal the number of rows of tbl or X.
Data Types: logical | single | double
Predictor data, specified as a numeric matrix.
Each row of X corresponds to one observation, and each column corresponds to one variable. The number of rows in X must equal the number of rows in Y.
The variables that make up the columns of X must have the same order as the predictor variables used to train mdl.
Data Types: double | single
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: loss(mdl,X,LossFun="exponential",UseParallel=true) specifies
to use an exponential loss function, and to run in parallel.
Loss function, specified as a built-in loss function name or a function handle.
The following table describes the values for the built-in loss functions.
| Value | Description |
|---|---|
"binodeviance" | Binomial deviance |
"classifcost" | Observed misclassification cost |
"classiferror" | Misclassified rate in decimal |
"exponential" | Exponential loss |
"hinge" | Hinge loss |
"logit" | Logistic loss |
"mincost" | Minimal expected misclassification cost (for classification scores that are posterior probabilities) |
"quadratic" | Quadratic loss |
"mincost"is appropriate for classification scores that are posterior probabilities.Bagged and subspace ensembles return posterior probabilities by default (
ens.Methodis"Bag"or"Subspace").To use posterior probabilities as classification scores when the ensemble method is
"AdaBoostM1","AdaBoostM2","GentleBoost", or"LogitBoost", you must specify the double-logit score transform by entering the following:ens.ScoreTransform = "doublelogit";For all other ensemble methods, the software does not support posterior probabilities as classification scores.
You can specify your own function using function handle notation. Suppose that
n is the number of observations in X, and
K is the number of distinct classes
(numel(mdl.ClassNames), where mdl is the input
model). Your function must have the signature
lossvalue = lossfun(C,S,W,Cost)The output argument
lossvalueis a scalar.You specify the function name (
lossfun).Cis ann-by-Klogical matrix with rows indicating the class to which the corresponding observation belongs. The column order corresponds to the class order inmdl.ClassNames.Create
Cby settingC(p,q) = 1, if observationpis in classq, for each row. Set all other elements of rowpto0.Sis ann-by-Knumeric matrix of classification scores. The column order corresponds to the class order inmdl.ClassNames.Sis a matrix of classification scores, similar to the output ofpredict.Wis ann-by-1 numeric vector of observation weights. If you passW, the software normalizes the weights to sum to1.Costis a K-by-Knumeric matrix of misclassification costs. For example,Cost = ones(K) - eye(K)specifies a cost of0for correct classification and1for misclassification.
For more details on loss functions, see Classification Loss.
Example: LossFun="binodeviance"
Example: LossFun=@Lossfun
Data Types: char | string | function_handle
Flag to run in parallel, specified as a numeric or logical
1 (true) or 0
(false). If you specify UseParallel=true, the
loss function executes for-loop iterations by
using parfor. The loop runs in parallel when you
have Parallel Computing Toolbox™.
Example: UseParallel=true
Data Types: logical
Observation weights, specified as a numeric vector or the name of a variable in
tbl. If you supply weights, loss
normalizes them so that the observation weights in each class sum to the prior
probability of that class.
If you specify Weights as a numeric vector, the size of
Weights must be equal to the number of observations in
X or tbl. The software normalizes
Weights to sum up to the value of the prior probability in the
respective class.
If you specify Weights as the name of a variable in
tbl, you must specify it as a character vector or string
scalar. For example, if the weights are stored as tbl.W, then
specify Weights as "W". Otherwise, the
software treats all columns of tbl, including
tbl.W, as predictors.
Example: Weights="W"
Data Types: single | double | char | string
Output Arguments
Classification loss, returned as a numeric scalar. L is a
scalar value, the loss for the entire ensemble of trees.
When computing the loss, the loss function
normalizes the class probabilities in ResponseVarName or
Y to the class probabilities used for training, which are stored
in the Prior property of mdl.
More About
Classification loss functions measure the predictive inaccuracy of classification models. When you compare the same type of loss among many models, a lower loss indicates a better predictive model.
Consider the following scenario.
L is the weighted average classification loss.
n is the sample size.
For binary classification:
yj is the observed class label. The software codes it as –1 or 1, indicating the negative or positive class (or the first or second class in the
ClassNamesproperty), respectively.f(Xj) is the positive-class classification score for observation (row) j of the predictor data X.
mj = yjf(Xj) is the classification score for classifying observation j into the class corresponding to yj. Positive values of mj indicate correct classification and do not contribute much to the average loss. Negative values of mj indicate incorrect classification and contribute significantly to the average loss.
For algorithms that support multiclass classification (that is, K ≥ 3):
yj* is a vector of K – 1 zeros, with 1 in the position corresponding to the true, observed class yj. For example, if the true class of the second observation is the third class and K = 4, then y2* = [
0 0 1 0]′. The order of the classes corresponds to the order in theClassNamesproperty of the input model.f(Xj) is the length K vector of class scores for observation j of the predictor data X. The order of the scores corresponds to the order of the classes in the
ClassNamesproperty of the input model.mj = yj*′f(Xj). Therefore, mj is the scalar classification score that the model predicts for the true, observed class.
The weight for observation j is wj. The software normalizes the observation weights so that they sum to the corresponding prior class probability stored in the
Priorproperty. Therefore,
Given this scenario, the following table describes the supported loss functions that you can specify by using the LossFun name-value argument.
| Loss Function | Value of LossFun | Equation |
|---|---|---|
| Binomial deviance | "binodeviance" | |
| Observed misclassification cost | "classifcost" | where is the class label corresponding to the class with the maximal score, and is the user-specified cost of classifying an observation into class when its true class is yj. |
| Misclassified rate in decimal | "classiferror" | where I{·} is the indicator function. |
| Cross-entropy loss | "crossentropy" |
The weighted cross-entropy loss is where the weights are normalized to sum to n instead of 1. |
| Exponential loss | "exponential" | |
| Hinge loss | "hinge" | |
| Logistic loss | "logit" | |
| Minimal expected misclassification cost | "mincost" |
The software computes the weighted minimal expected classification cost using this procedure for observations j = 1,...,n.
The weighted average of the minimal expected misclassification cost loss is |
| Quadratic loss | "quadratic" |
If you use the default cost matrix (whose element value is 0 for correct classification
and 1 for incorrect classification), then the loss values for
"classifcost", "classiferror", and
"mincost" are identical. For a model with a nondefault cost matrix,
the "classifcost" loss is equivalent to the "mincost"
loss most of the time. These losses can be different if prediction into the class with
maximal posterior probability is different from prediction into the class with minimal
expected cost. Note that "mincost" is appropriate only if classification
scores are posterior probabilities.
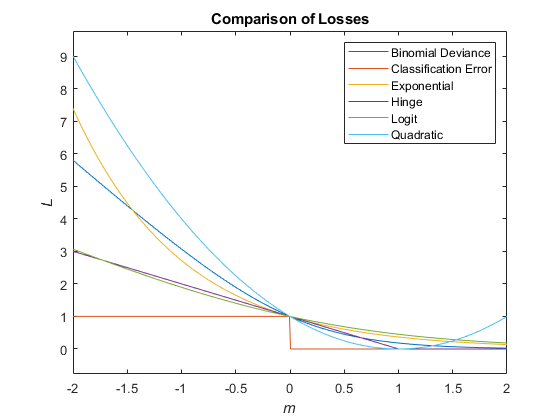
This figure compares the loss functions (except "classifcost",
"crossentropy", and "mincost") over the score
m for one observation. Some functions are normalized to pass through
the point (0,1).
Extended Capabilities
Usage notes and limitations:
You cannot use the
UseParallelname-value argument with tall arrays.
For more information, see Tall Arrays.
To run in parallel, set the UseParallel name-value argument to
true in the call to this function.
For more general information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
You cannot use the UseParallel name-value
argument with tall arrays, GPU arrays, or code generation.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2026a
See Also
margin | edge | predict | CompactClassificationXGBoost | importModelFromXGBoost
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)