mvncdf
Multivariate normal cumulative distribution function
Syntax
Description
p = mvncdf(X)X. For more information, see Multivariate Normal Distribution.
p = mvncdf(___,options)p, using any of the input argument combinations in the
previous syntaxes. Create the options argument using the
statset function with any
combination of the parameters 'TolFun',
'MaxFunEvals', and 'Display'.
[
additionally returns an estimate of the error in p,err] = mvncdf(___)p. For
more information, see Algorithms.
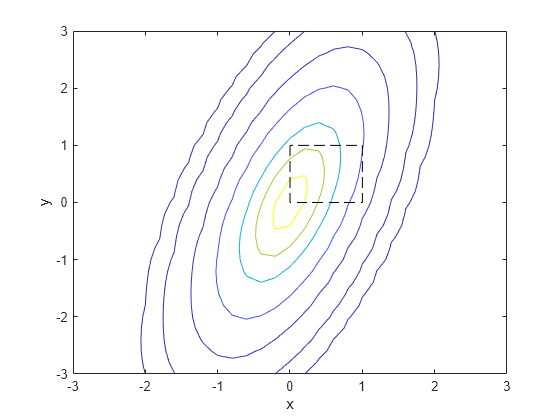
Examples
Input Arguments
Output Arguments
More About
Tips
In the one-dimensional case,
Sigmais the variance, not the standard deviation. For example,mvncdf(1,0,4)is the same asnormcdf(1,0,2), where4is the variance and2is the standard deviation.
Algorithms
For bivariate and trivariate distributions, mvncdf uses adaptive
quadrature on a transformation of the t density, based on methods
developed by Drezner and Wesolowsky [1]
[2]
and by Genz [3]. For four or more dimensions,
mvncdf uses a quasi-Monte Carlo integration algorithm based on
methods developed by Genz and Bretz [4]
[5].
References
[1] Drezner, Z. “Computation of the Trivariate Normal Integral.” Mathematics of Computation. Vol. 63, 1994, pp. 289–294.
[2] Drezner, Z., and G. O. Wesolowsky. “On the Computation of the Bivariate Normal Integral.” Journal of Statistical Computation and Simulation. Vol. 35, 1989, pp. 101–107.
[3] Genz, A. “Numerical Computation of Rectangular Bivariate and Trivariate Normal and t Probabilities.” Statistics and Computing. Vol. 14, No. 3, 2004, pp. 251–260.
[4] Genz, A., and F. Bretz. “Numerical Computation of Multivariate t Probabilities with Application to Power Calculation of Multiple Contrasts.” Journal of Statistical Computation and Simulation. Vol. 63, 1999, pp. 361–378.
[5] Genz, A., and F. Bretz. “Comparison of Methods for the Computation of Multivariate t Probabilities.” Journal of Computational and Graphical Statistics. Vol. 11, No. 4, 2002, pp. 950–971.
[6] Kotz, S., N. Balakrishnan, and N. L. Johnson. Continuous Multivariate Distributions: Volume 1: Models and Applications. 2nd ed. New York: John Wiley & Sons, Inc., 2000.
Extended Capabilities
Version History
Introduced in R2006a