umap
Uniform Manifold Approximation and Projection (UMAP) for dimension reduction
Since R2026a
Description
Y = umap(X)X.
To calculate the embeddings, the function uses the Neg-t-SNE version of the Uniform Manifold
Approximation and Projection (UMAP) algorithm for dimension reduction. For more information,
see Algorithms.
Y = umap(X,Name=Value)
[
also returns the Y,NeighborIndicesResult] = umap(___)NumNeighbors nearest neighbor row indices for each row
of X that does not contain any NaN values. Use this
syntax with any of the input arguments in the previous syntaxes.
Examples
View the 2-D and 3-D embeddings of the human activity data set using the umap function.
Load the data set.
load humanactivityThe data set contains 24,075 observations of 60 predictors, and an activity class label for each observation. For more details on the data set, enter Description at the command line.
Associate the activities with the labels in actid.
activities = ["Sitting";"Standing";"Walking";"Running";"Dancing"]; activity = activities(actid);
Because UMAP is a stochastic algorithm, set the random number seed.
rng(0,"twister") % For reproducibility
View the 2-D embedding. Assign a color to each activity class using the hsv colormap.
Y2 = umap(feat); % Default number of embedding dimensions is 2 figure colormap = hsv(5); gscatter(Y2(:,1),Y2(:,2),activity,colormap) title("2-D Embedding")
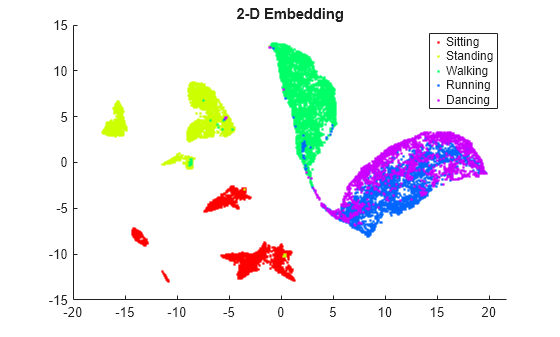
View the 3-D embedding.
Y3= umap(feat,NumDimensions=3); figure scatter3(Y3(:,1),Y3(:,2),Y3(:,3),8,colormap(actid,:),"filled") title("3-D Embedding") grid on view([12 60])
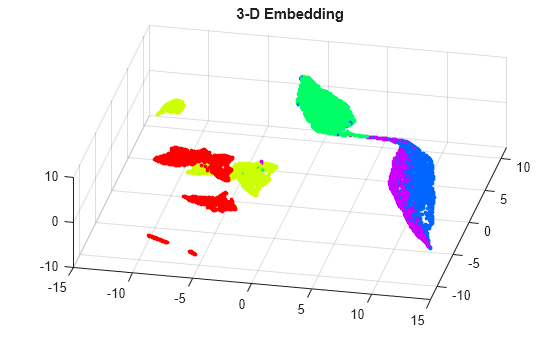
By rotating the 3-D plot, you can see that the activities Running and Dancing are more easily distinguished in 3-D than in 2-D.
Determine the effects of the UMAP embedding density and number of nearest neighbor settings on the two-dimensional embeddings of two data sets: a simulated clustered data set and a human activity data set.
Create Simulated Clustered Data
Create a simulated clustered data set with 1000 observations of 10 predictors. X contains five clusters of 200 observations each. Y contains the cluster identification numbers. The predictor values of each cluster centroid lie within the range [–5,5] and have a standard deviation sigma. The sigma value for each cluster is a random scalar in the range (0,3].
rng(0,"twister"); % For reproducibility nClusters = 5; obsPerCluster = 200; X = []; Y = []; xrange = 5; nPredictors = 10; sigmaRange = 3; for c = 1:nClusters Y = [Y; c*ones(obsPerCluster,1)]; sigma = rand*sigmaRange; X = [X; randn(obsPerCluster,nPredictors)*sigma + ... (randi(2*xrange,[1,nPredictors])-xrange).* ... ones(obsPerCluster,nPredictors)]; end
View Two-Dimensional Clustered Data Embeddings
Compute two-dimensional embeddings for X with different parameter settings by using the umap function. Specify 0.1, 1, and 10 for the embedding density values, and 3, 15, and 100 for the number of nearest neighbor values. Because UMAP is a stochastic algorithm, reset the random number seed each time and set Reproducible to true. This setting for Reproducible slows down computations considerably, but is needed in this example to ensure a fair comparison between parameter settings. Display each embedding in a separate plot and assign a different color to each cluster identification number.
density = [0.1 1 10 0.1 1 10 0.1 1 10]; nNeighbors = [3 3 3 15 15 15 100 100 100]; figure t = tiledlayout(3,3,TileSpacing="compact",Padding="compact"); for i = 1:9 rng(0,"twister"); % For fair comparison E1 = umap(X,EmbeddingDensity=density(i), ... NumNeighbors=nNeighbors(i),Reproducible=false); ax = nexttile; gscatter(E1(:,1),E1(:,2),Y,[],"o",3,"off") ax.XTickLabel = []; ax.YTickLabel = []; if i > 6 xlabel(sprintf("EmbeddingDensity = %.1f",density(i)),FontSize=8); end if (mod(i-1,3) == 0) ylabel(sprintf("NumNeighbors = %d",nNeighbors(i)), ... FontSize=8,Rotation=0); end axis square end
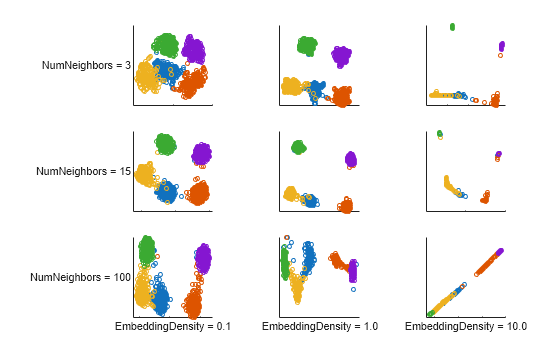
Load and Preprocess Human Activity Data
Load the human activity data set.
load humanactivityThe data set contains 24,075 observations of 60 predictors, and an activity class label for each observation. For more details on the data set, enter Description at the command line.
The observations are organized by activity class. To better represent a random set of data, shuffle the rows.
n = numel(actid); idx = randsample(n,n); X2 = feat(idx,:); actid = actid(idx);
View Two-Dimensional Human Activity Data Embeddings
Compute two-dimensional embeddings using standardized data and the same set of embedding density values and number of nearest neighbor values as for the simulated clustered data set. Reset the random number seed before computing each embedding, and set Reproducible=true to ensure a fair comparison between parameter settings. Display each embedding in a separate plot, and assign a different color to each activity class.
figure t = tiledlayout(3,3,TileSpacing="compact",Padding= "compact"); for i = 1:9 rng(0,"twister"); % For fair comparison E2 = umap(X2,Standardize=true,EmbeddingDensity=density(i), ... NumNeighbors=nNeighbors(i),Reproducible=true); ax = nexttile; gscatter(E2(:,1),E2(:,2),actnames(actid)',[],"o",3,"off") ax.XTickLabel = []; ax.YTickLabel = []; if i > 6 xlabel(sprintf("EmbeddingDensity = %.1f",density(i)),FontSize=8); end if (mod(i-1,3) == 0) ylabel(sprintf("NumNeighbors = %d",nNeighbors(i)), ... FontSize=8,Rotation=0); end axis square end
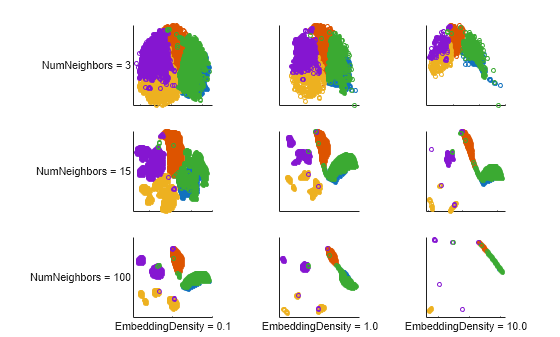
The embeddings of the two data sets show that higher embedding density values result in a tighter clustering of points. For very high values of EmbeddingDensity, the clusters can begin to merge. The effect of the number of nearest neighbors parameter depends on the particular data set and the embedding density value. A higher value of NumNeighbors causes umap to treat more pairs of points as neighbors, and to try bringing them closer together in the embedding space.
For the simulated clustered data set, the tightest, most distinct cluster representation occurs for NumNeighbors = 15. However, when EmbeddingDensity is 10, none of the NumNeighbors values yield an embedding with distinct clusters.
For the human activity data set, the highest NumNeighbors value yields relatively distinct clusters. However, when NumNeighbors is 100 and EmbeddingDensity is 10, most of the observations overlap in the same locations.
Input Arguments
Data points, specified as a numeric matrix with m columns, or as
a table with m variables. Each row of X contains
one m-dimensional point. X must have at least
two rows that do not contain a NaN value. umap
ignores rows of X that contain at least one NaN
value.
Data Types: single | double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: Y = umap(X,NumDimensions=3) computes a three-dimensional
embedding for the data points X.
Algorithm Control
Size of the Gram matrix in megabytes, specified as a positive scalar or
"maximal". This argument is valid only when the
Distance name-value argument begins with
fast.
If you set CacheSize to "maximal",
umap tries to allocate enough memory for an entire
intermediate matrix whose size is M-by-M, where
M is the number of rows of the input data X.
The cache size does not have to be large enough for an entire intermediate matrix, but
must be at least large enough to hold an M-by-1 vector. Otherwise,
umap uses the standard algorithm for computing Euclidean
distances.
If the value of CacheSize is too large or
"maximal", umap might try to allocate
a Gram matrix that exceeds the available memory. In this case, the software issues an
error.
Example: CacheSize="maximal"
Data Types: double | char | string
Distance metric, specified as a character vector, string scalar, or function handle, as described in the following table.
| Value | Description |
|---|---|
"euclidean" | Euclidean distance (default) |
"seuclidean" | Standardized Euclidean distance. Each coordinate difference between the
rows in X and the query matrix is scaled by dividing by the
corresponding element of the standard deviation computed from
S = std(X,"omitnan"). |
"fasteuclidean" | Euclidean distance computed by using an alternative algorithm that saves
time when the number of columns in X is at least 10. In
some cases, this faster algorithm can reduce accuracy. Algorithms starting
with "fast" do not support sparse data. For details, see
Algorithms. |
"fastseuclidean" | Standardized Euclidean distance computed by using an alternative
algorithm that saves time when the number of columns in X
is at least 10. In some cases, this faster algorithm can reduce accuracy.
Algorithms starting with "fast" do not support sparse data.
For details, see Algorithms. |
"mahalanobis" | Mahalanobis distance, computed using the positive definite
covariance matrix |
"cityblock" | City block distance |
"minkowski" | Minkowski distance with exponent 2. This distance is the same as the Euclidean distance. |
"chebychev" | Chebychev distance, which is the maximum coordinate difference |
"cosine" | One minus the cosine of the included angle between observations (treated as vectors) |
"correlation" | One minus the sample linear correlation between observations (treated as sequences of values) |
"hamming" | Hamming distance, which is the percentage of coordinates that differ |
"jaccard" | One minus the Jaccard coefficient, which is the percentage of nonzero coordinates that differ |
"spearman" | One minus the sample Spearman's rank correlation between observations (treated as sequences of values) |
@ | Custom distance function handle. A distance function has the form function D2 = distfun(ZI,ZJ) % calculation of distance ...
If your data is not sparse, you can generally compute distances more quickly by using a built-in distance metric instead of a function handle. |
In all cases, umap uses squared pairwise distances to
calculate the Cauchy kernel in the joint distribution of
X.
Example: Distance="mahalanobis"
Data Types: char | string | function_handle
Factor for adjusting the density of the embedding structure, specified as a
positive scalar. A larger EmbeddingDensity value increases the
attraction between similar points, resulting in a more continuous embedding with
densely packed regions. A smaller EmbeddingDensity value
increases the repulsion, leading to a more discrete embedding structure where the
points are more widely spread apart.
This argument corresponds to the parameter in the Neg-t-SNE algorithm of Damrich et al. [4]. The default value
(1) corresponds approximately to the behavior of the original
UMAP algorithm of McInnes et al. [6].
Example: EmbeddingDensity=10
Data Types: single | double
Initialization method for the low-dimensional embedding, specified as
"pca" or "random". The initial embedding is an
n-by-NumDimensions matrix, where
n is the number of rows of X that do not
contain any NaN values.
"pca"— Use principal component analysis (PCA) to initialize the embedding. Before embedding the high-dimensional data,umapfirst reduces the dimensionality of the data toNumDimensionsPCA components by using thepcafunction. Compared to using random initial positions, this method is generally better at preserving the global structure of the high-dimensional data."random"— Initialize the embedding using randomly assigned positions.
Example: Initialization="random"
Data Types: char | string
Number of nearest neighbors, specified as a positive integer. A higher value of
NumNeighbors causes umap to treat more
pairs of points as neighbors, and to try to bring them closer together in the
embedding space.
If you specify NeighborIndices, then the default value of
NumNeighbors is n, where
n is the number of columns in
NeighborIndices. If you specify both
NeighborIndices and NumNeighbors, then
NumNeighbors must equal n.
Example: NumNeighbors=10
Data Types: single | double
Precomputed nearest neighbor indices, specified as a matrix of positive integers.
Setting this name-value argument can improve performance by preventing redundant
computations. Each row of NeighborIndices contains the indices of
the nearest neighbors for the corresponding observation (row) in
X (after the function removes any rows in
X that contain at least one NaN value). If
you specify NumNeighbors, then
NeighborIndices must contain NumNeighbors
columns.
Data Types: single | double
Flag to enforce reproducibility, specified as "on" or
"off", or as a logical 0
(false) or 1 (true). To
reproduce the embeddings over repeated runs, specify Reproducible
as true and set the seed of the random number generator using
rng.
Note
Setting Reproducible to true can cause
slower embedding computation times.
Example: Reproducible=true
Data Types: single | double | logical
Flag to normalize the input data, specified as "on" or
"off", or as a logical 0
(false) or 1 (true). When
the value is true, umap centers and
scales each column of X by first subtracting its mean, then
dividing by its standard deviation.
When features in X are on different scales, set
Standardize to true. The learning process is
based on nearest neighbors, so features with large scales might override the
contribution of features with small scales.
Example: Standardize=true
Data Types: single | double | logical
Optimization Control
Number of optimization epochs, specified as a positive integer. A smaller
NumEpochs value speeds up computation time, but can lead to a
poorly optimized embedding.
Example: NumEpochs=100
Data Types: single | double
Learning rate for the optimization process, specified as a positive scalar.
When LearnRate is too small, umap can
converge to a poor local minimum. When LearnRate is too large, the
algorithm might not achieve the best possible optimization for the specified value of
NumEpochs.
Example: LearnRate=3
Data Types: single | double
Output Arguments
Embedded points, returned as an
n-by-NumDimensions matrix. Each row represents
one embedded point. n is the number of rows of X
that do not contain any NaN values.
Nearest neighbors for the data points in X, returned as an
n-by-NumNeighbors matrix of row indices, where
n is the number of rows of X that do not
contain any NaN values.
Algorithms
The UMAP algorithm creates a set of embedded points in a low-dimensional space whose relative similarities mimic those of the original high-dimensional points. The embedded points reflect the clustering in the original data. Unlike PCA, which uses a well-quantified linear algorithm, UMAP provides a nonlinear representation of high-dimensional data. This representation can have an arbitrary rotation or mirroring. However, UMAP can capture high-dimensional topological structures in the data that cannot be represented by principal components.
Because the stochastic nature of the UMAP algorithm makes the embedding sensitive to
parameters such as the number of nearest neighbors, embedding density, learning rate, number
of optimization epochs, and initialization, no "true" embedding solution exists for any
particular data set (see UMAP Settings). The default values of
these and other parameters are only suggested starting points. A best practice is to run the
umap function multiple times using different parameter settings and
compare the embeddings to those generated by other algorithms, such as tsne. In general, consider using umap and
tsne only as a starting point for further experiments and hypotheses
regarding the data.
The umap function uses the Neg-t-SNE version of UMAP developed by
Damrich et al. [4]. The Neg-t-SNE algorithm
primarily differs from the original UMAP algorithm of McInnes et al. [6] by using a loss function
that is less vulnerable to numerical instability, and by incorporating a normalization
parameter (see EmbeddingDensity) that controls whether the output
embedding is more similar to t-SNE or the original UMAP. During optimization of the embedding,
the algorithm attempts to pull together pairs of positive samples, which
are close together in the high-dimensional space, and push apart negative
samples, which are not close together in the high-dimensional space. For each
positive pair, umap samples a fixed number of negative samples
(m = 5). As a result, the embedding computation times for UMAP are
typically much faster than those for t-SNE, making UMAP more suitable for large data sets and
for investigating the effects of different parameter settings on the embedding.
The Distance argument values that begin with fast
(such as "fasteuclidean" and "fastseuclidean")
calculate Euclidean distances using an algorithm that uses extra memory to save
computational time. This algorithm is named "Euclidean Distance Matrix Trick" in Albanie
[1] and elsewhere. Internal
testing shows that this algorithm saves time when the number of predictors is at least 10.
Algorithms starting with fast do not support sparse data.
To find the matrix D of distances between all the points xi and xj, where each xi has n variables, the algorithm computes distance using the final line in the following equations:
The matrix in the last line of the equations is called the Gram matrix. Computing the set of squared distances is faster, but slightly less numerically stable, when you compute and use the Gram matrix instead of computing the squared distances by squaring and summing. For more details, see Albanie [1].
To store the Gram matrix, the software uses a cache with the default size of
1e3 megabytes. You can set the cache size using the
CacheSize name-value argument. If the value of
CacheSize is too large or "maximal", then the
software might try to allocate a Gram matrix that exceeds the available memory. In this
case, the software issues an error.
References
[1] Albanie, Samuel. Euclidean Distance Matrix Trick. June, 2019. Available at https://samuelalbanie.com/files/Euclidean_distance_trick.pdf.
[2] Böhm, J. N. "Attraction-Repulsion Spectrum in Neighbor Embeddings." Journal of Machine Learning Research , no. 23 (2022): 4118–4149.
[3] Damrich, Sebastian, and Fred A. Hamprecht. "On UMAP's True Loss Function." Neural Information Processing Systems (2021).
[4] Damrich, Sebastian, et al. "From t-SNE to UMAP with Contrastive Learning." arXiv:2206.01816 [cs], June 2022. arXiv.org.
[5] Healy, John, and Leland McInnes. "Uniform Manifold Approximation and Projection." Nature Reviews Methods Primers 4, no. 1 (2024).
[6] McInnes, Leland, John Healy, Nathaniel Saul, and Lukas Großberger. "UMAP: Uniform Manifold Approximation and Projection." Journal of Open Source Software 3, no. 29 (September 2, 2018): 861.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)