knnsearch
使用输入数据查找 k 最近邻点
说明
示例
输入参数
名称-值参数
输出参量
提示
对于固定正整数 k,
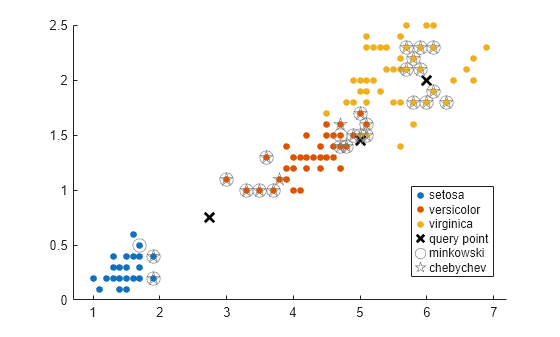
knnsearch在X中找到 k 个最接近Y中每个点的点。要找到X中在固定距离内接近Y中每个点的所有点,请使用rangesearch。knnsearch不保存搜索对象。要创建一个搜索对象,请使用createns。
算法
替代功能
如果您将 knnsearch 函数的 'NSMethod' 名称-值对组参量设置为合适的值('exhaustive' 用于穷举搜索算法,或 'kdtree' 用于 Kd 树算法),则搜索结果等效于使用 knnsearch 对象函数进行距离搜索得到的结果。与 knnsearch 函数不同,knnsearch 对象函数需要 ExhaustiveSearcher 或 KDTreeSearcher 模型对象。
Simulink 模块
要将 k 最近邻搜索集成到 Simulink® 中,您可以使用 Statistics and Machine Learning Toolbox™ 库中的 KNN Search 模块或具有 knnsearch 函数的 MATLAB Function 模块。有关示例,请参阅Predict Class Labels Using MATLAB Function Block。
在决定要使用的方法时,请考虑以下因素:
如果使用 Statistics and Machine Learning Toolbox 库模块,可以使用定点工具 (Fixed-Point Designer)将浮点模型转换为定点。
必须为具有
knnsearch函数的 MATLAB Function 模块启用对可变大小数组的支持。
参考
[1] Friedman, J. H., J. Bentley, and R. A. Finkel. “An Algorithm for Finding Best Matches in Logarithmic Expected Time.” ACM Transactions on Mathematical Software 3, no. 3 (1977): 209–226.