waveletPooling2dLayer
Description
A 2-D discrete wavelet pooling layer applies the forward and inverse discrete wavelet transforms to reconstruct approximations of the layer input. Use this layer to downsample the layer input along the spatial dimensions. The layer supports learnable (adaptive) and nonadaptive pooling. For more information, see Discrete Wavelet Pooling. Use of this layer requires Deep Learning Toolbox™.
Creation
Description
layer = waveletPooling2dLayer
The input to waveletPooling2dLayer must be a real-valued dlarray (Deep Learning Toolbox) object in
"SSCB" format. The output is a dlarray object in
"SSCB" format. For more information, see Layer Output Format.
Note
When you initialize the learnable parameters of waveletPooling2dLayer, the
layer weights are set to unity. It is not recommended to initialize the weights
directly.
layer = waveletPooling2dLayer(PropertyName=Value)
Example: layer =
waveletPooling2dLayer(Wavelet="db4",Boundary="zeropad") creates a wavelet
pooling layer that uses the extremal phase Daubechies wavelet with four vanishing moments
and zero padding at the boundaries.
Properties
DWT
This property is read-only after object creation.
Real-valued orthogonal or biorthogonal wavelet, specified as a string scalar or
character vector recognized by wavemngr. The layer uses the specified wavelet in the 2-D DWT.
Data Types: char | string
This property is read-only after object creation.
Reconstruction level, specified as a nonnegative integer between
0 and min(log2([size(X,1) size(X,2)])), where
X is the layer input. The layer reconstructs the pooled output at
level ReconstructionLevel. By default, the layer output is a
factor-of-2 dimension reduction for each input feature.
If you set ReconstructionLevel, then the default value of
AnalysisLevel is ReconstructionLevel+1.
For more information about the size of the layer output, see Layer Output Format.
Data Types: single | double
This property is read-only after object creation.
Analysis level, or decomposition level, of the 2-D DWT, specified as a positive
integer between 1 and min(log2([size(X,1)
size(X,2)])), where X is the layer input.
AnalysisLevel must be greater than
ReconstructionLevel.
If you set AnalysisLevel, then the default value of
ReconstructionLevel is AnalysisLevel−1.
Data Types: single | double
This property is read-only after object creation.
Boundary extension mode to use in the DWT, specified as
"reflection", "periodic", or
"zeropad". The layer extends the coefficients at the boundary at
each decomposition level based on the corresponding mode in dwtmode:
"reflection"— Half-point symmetric extension,"sym""periodic"— Periodic extension,"per""zeropad— Zero padding,"zpd"
To learn how the boundary extension mode can affect the size of the layer output, see Layer Output Format.
This property is read-only after object creation.
Mask indicating which detail (wavelet) coefficients to include in pooling, specified as a logical array.
The shape of the array is
Nchannel-by-3-by-Ndiff, where
Nchannel is the number of channels in the layer input, 3
represents the three wavelet subbands in the order LH (horizontal details), HL
(vertical details), and HH (diagonal details), and Ndiff is the
difference between the analysis level and reconstruction level. The pages of the array
correspond to the decomposition levels ordered from finer to coarser scale. The layer
includes and excludes the detail coefficients for pooling when the corresponding value
in the mask is 1 (true) or 0
(false), respectively.
If
Nchannelis equal to1and the input to the layer's forward / predict object function has more than one channel, the layer expands the values ofSelectedDetailCoefficientsin the 1-by-3-by-Ndiffarray to all channels.If you set
Ndiffto1and the difference between the analysis and reconstruction levels is greater than 1, the layer expands theSelectedDetailCoefficientsvalues to all levels.
The following 1-D example illustrates how
SelectedDetailCoefficients interacts with
DetailWeights. The behavior is identical for the 2-D wavelet
pooling layer. If SelectedDetailCoefficients = [1 0
1]DetailWeights = [0.5 1e3
0.4]0, the coefficients at that
level are ignored. The gain corresponding to the middle level is effectively
zero.
Data Types: logical
This property is read-only after object creation.
Expected size of the layer output along the spatial dimensions, specified as
"none" or a two-element vector. By default, the output size is
not verified. If you set ExpectedOutputSize and the output sizes
do not match the specified value, the result is an error.
ExpectedOutputSize plays an analogous role to the bookkeeping
matrix S output argument of wavedec2, when that information is missing. The bookkeeping matrix
contains the dimensions of the coefficients by level and the dimensions of the
input.
Parameters
Detail (wavelet) coefficient subband weights, specified as [],
a numeric array, or a dlarray object.
These layer weights are learnable parameters. DetailWeights
is a Nchannel-by-3-by-Ndiff tensor, where
Nchannel is the number of channels in the layer input, 3
represents the three wavelet subbands in the order LH (horizontal details), HL
(vertical details), and HH (diagonal details), and Ndiff is the
difference between the analysis level and reconstruction level. You can use the
function initialize (Deep Learning Toolbox)
to initialize the learnable parameters of a deep learning neural network that includes
waveletPooling2dLayer objects. When you initialize the layers,
initialize sets DetailWeights to
ones(Nchannel,3,.AnalysisLevel-ReconstructionLevel)
You must initialize waveletPooling2dLayer before using the layer's
forward/predict method.
Data Types: single | double
Lowpass (scaling) coefficient subband weights, specified as [],
a numeric array, or a dlarray object.
These layer weights are learnable parameters. LowpassWeights
is a Nchannel-byNdiff tensor, where
Nchannel is the number of channels in the layer input and
Ndiff is the difference between the analysis level and
reconstruction level. You can use the function initialize (Deep Learning Toolbox)
to initialize the learnable parameters of a deep learning neural network that includes
waveletPooling2dLayer objects. When you initialize the layers,
initialize sets LowpassWeights to
ones(Nchannel,.AnalysisLevel-ReconstructionLevel)
You must initialize waveletPooling2dLayer before using the layer's
forward/predict method.
It is not recommended to initialize the weights directly.
Data Types: single | double
Learning Rate
Learning rate factor for the detail (wavelet) coefficient subband weights,
specified as a nonnegative scalar. By default, the weights do not update with
training. You can also set this property using the function setLearnRateFactor (Deep Learning Toolbox).
To perform adaptive pooling, specify a nonzero learning rate factor. When
DetailWeightLearnRateFactor is nonzero, the layer uses the
detail subband weights to modify the wavelet coefficients.
Data Types: single | double
Learning rate factor for the lowpass (LL) coefficient subband weights, specified
as a nonnegative scalar. By default, the weights do not update with training. You can
also set this property using the function setLearnRateFactor (Deep Learning Toolbox).
To perform adaptive pooling, specify a nonzero learning rate factor. When
LowpassWeightLearnRateFactor is nonzero, the layer uses the
lowpass weights to modify the lowpass (scaling) coefficients.
Data Types: single | double
Layer
This property is read-only.
Number of inputs to the layer, stored as 1. This layer accepts a
single input only.
Data Types: double
This property is read-only.
Input names, stored as {'in'}. This layer accepts a single input
only.
Data Types: cell
This property is read-only.
Number of outputs from the layer, stored as 1. This layer has a
single output only.
Data Types: double
This property is read-only.
Output names, stored as {'out'}. This layer has a single output
only.
Data Types: cell
Examples
Load the xbox image. The image is a 128-by-128 matrix. Save the image in single precision as a dlarray object with format "SSCB".
load xbox dlimg = dlarray(single(xbox),"SSCB");
Create the default 2-D discrete wavelet pooling layer. By default, the learnable parameters DetailWeights and LowpassWeights are both empty.
dwtPool = waveletPooling2dLayer
dwtPool =
waveletPooling2dLayer with properties:
Name: ''
DetailWeightLearnRateFactor: 0
LowpassWeightLearnRateFactor: 0
Wavelet: 'db1'
ReconstructionLevel: 1
AnalysisLevel: 2
Boundary: 'reflection'
SelectedDetailCoefficients: [1 1 1]
ExpectedOutputSize: 'none'
Learnable Parameters
DetailWeights: []
LowpassWeights: []
State Parameters
No properties.
Show all properties
Include the default 2-D wavelet pooling layer in a Layer array.
layers = [ ...
imageInputLayer([128 128 1])
waveletPooling2dLayer]layers =
2×1 Layer array with layers:
1 '' Image Input 128×128×1 images with 'zerocenter' normalization
2 '' waveletPooling2dLayer waveletPooling2dLayer
Convert the layer array to a dlnetwork object. Because the layer array has an input layer and no other inputs, the software initializes the network.
dlnet = dlnetwork(layers)
dlnet =
dlnetwork with properties:
Layers: [2×1 nnet.cnn.layer.Layer]
Connections: [1×2 table]
Learnables: [2×3 table]
State: [0×3 table]
InputNames: {'imageinput'}
OutputNames: {'layer'}
Initialized: 1
View summary with summary.
Confirm the network learnable parameters are DetailWeights and LowpassWeights.
dlnet.Learnables
ans=2×3 table
Layer Parameter Value
_______ ________________ _____________
"layer" "DetailWeights" {1×3 dlarray}
"layer" "LowpassWeights" {1×1 dlarray}
Inspect the wavelet pooling layer in the network. Confirm the software initialized the weights.
dlnet.Layers(2)
ans =
waveletPooling2dLayer with properties:
Name: 'layer'
DetailWeightLearnRateFactor: 0
LowpassWeightLearnRateFactor: 0
Wavelet: 'db1'
ReconstructionLevel: 1
AnalysisLevel: 2
Boundary: 'reflection'
SelectedDetailCoefficients: [1 1 1]
ExpectedOutputSize: 'none'
Learnable Parameters
DetailWeights: [1×3 dlarray]
LowpassWeights: [1×1 dlarray]
State Parameters
No properties.
Show all properties
Run the image through the network. By default, the layer outputs the level-one lowpass approximation of the input.
dlnetout = forward(dlnet,dlimg); size(dlnetout)
ans = 1×4
64 64 1 1
dims(dlnetout)
ans = 'SSCB'
Plot the layer input and output.
dlnetout2 = squeeze(extractdata(dlnetout)); tiledlayout(1,2) nexttile imagesc(xbox) title("Layer Input") nexttile imagesc(dlnetout2) title("Layer Output") colormap gray
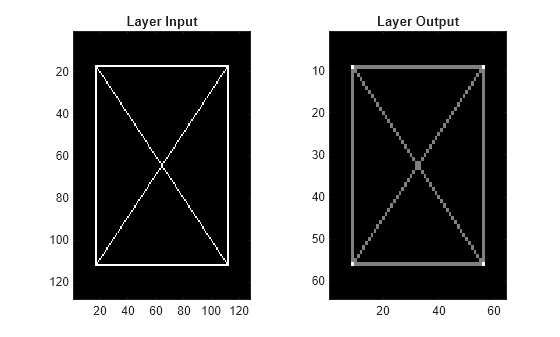
Load the xbox image. The image dimensions are 128-by-128. Save the image in single precision as a dlarray object with format "SSCB".
load xbox dlimg = dlarray(single(xbox),"SSCB");
Create a 2-D discrete wavelet pooling layer that uses the biorthogonal bior4.4 wavelet, which has four vanishing moments each for the decomposition and reconstruction filters. Set the analysis level to 4 and reconstruction level to 1. Set the boundary extension mode to "zeropad". Specify a mask so that the layer excludes from the pooling:
The LH subband (horizontal details) at level 3
The HL subband (vertical details) at level 2
The wavelet HH subband (diagonal details) at levels 3 and 4
% specify levels alevel = 4; rlevel = 1; bdy = "zeropad"; wv = "bior4.4"; % create mask msk = ones(1,3,alevel-rlevel); msk(1,1,2) = 0; % exclude LH subband msk(1,2,1) = 0; % exclude HL subband msk(1,3,2:3) = 0; % exclude HH subband % create layer poolDWT = waveletPooling2dLayer(AnalysisLevel=alevel, ... ReconstructionLevel=rlevel, ... SelectedDetailCoefficients=msk, ... Wavelet=wv, ... Boundary=bdy);
Create a Layer array containing an image input layer and the pooling layer.
layers = [ ...
imageInputLayer([size(dlimg,1:2) 1])
poolDWT];Convert the layer array to a dlnetwork object.
dlnet = dlnetwork(layers); dlnet.Layers(2)
ans =
waveletPooling2dLayer with properties:
Name: 'layer'
DetailWeightLearnRateFactor: 0
LowpassWeightLearnRateFactor: 0
Wavelet: "bior4.4"
ReconstructionLevel: 1
AnalysisLevel: 4
Boundary: 'zeropad'
SelectedDetailCoefficients: [1×3×3 double]
ExpectedOutputSize: 'none'
Learnable Parameters
DetailWeights: [1×3×3 dlarray]
LowpassWeights: [1×1 dlarray]
State Parameters
No properties.
Show all properties
Run the signal through the network.
netout = forward(dlnet,dlimg);
Now use the same layer property values in the functions dldwt and dlidwt.
Use the dldwt function to obtain the differentiable DWT of the image down to level 4. To specify the extension mode, use the PaddingMode name-value argument. To obtain the full wavelet decomposition instead of only the final level wavelet coefficients, set the full wavelet decomposition option to true.
[A,D] = dldwt(dlimg, ... Wavelet=wv, ... Level=alevel, ... FullTree=true, ... PaddingMode=bdy);
For a multilevel 2-D inverse DWT, the dlidwt function expects the wavelet subband gains (mask) to be an NC-by-3-by-L matrix, where NC is the number of channels in the data, and L is the difference between the decomposition level (used to obtain the coefficients) and the reconstruction level. Use the dlidwt function to reconstruct the DWT up to level 2. Set the extension mode to zero padding and the DetailGain name-value argument to the mask.
dlout = dlidwt(A,D, ... Wavelet=wv, ... Level=rlevel, ... DetailGain=msk, ... PaddingMode=bdy);
Confirm the sizes of the layer output and dlidwt output are identical.
size(netout)
ans = 1×4
68 68 1 1
size(dlout)
ans = 1×4
68 68 1 1
Confirm the outputs are equal.
netoutEx = extractdata(netout); dloutEx = extractdata(dlout); max(abs(netoutEx(:)-dloutEx(:)))
ans = single
0
Plot the original image and its pooled approximation.
tiledlayout(1,2) nexttile imagesc(xbox) axis tight title("Original Image") nexttile imagesc(netoutEx) axis tight title("Pooled Approximation") colormap gray
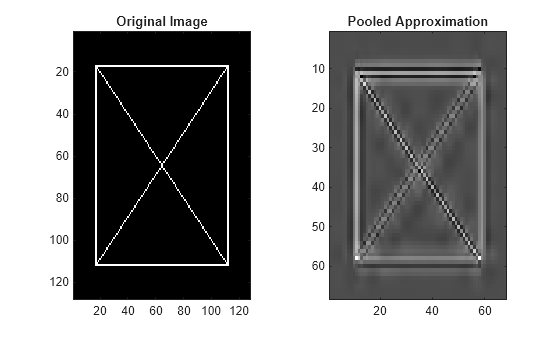
Create a 127-by-135 image. Save the image in single precision as a dlarray object.
img = randn(127,135);
dlimg = dlarray(img,"SSCB");The reconstruction level and the difference between the analysis and reconstruction levels can affect the output size.
Reconstruction Level Greater Than 0, Difference Between Analysis and Reconstruction Levels Greater Than 1
Use the dldwt function to obtain the differentiable DWT of the image down to level 4. Specify the db2 wavelet and zero padding as the extension mode. Obtain the full wavelet decomposition by setting FullTree to true.
wv = "db2"; alevel = 4; bdy = "zeropad"; [A,D] = dldwt(dlimg, ... Wavelet=wv, ... Level=alevel, ... FullTree=true, ... PaddingMode=bdy); %#ok<*ASGLU>
Create a waveletPooling2dLayer using the same parameters as the dldwt function. Specify a reconstruction level of 2.
rlevel = 2; wpl = waveletPooling2dLayer(Wavelet=wv, ... AnalysisLevel=alevel, ... ReconstructionLevel=rlevel, ... Boundary=bdy);
Create a Layer array containing an image input layer appropriate for the signal and the wavelet pooling layer. Convert the layer array to a dlnetwork object. Run the image through the network.
layers = [
imageInputLayer([size(dlimg,1:2) 1])
wpl];
dlnet = dlnetwork(layers);
netOutput = forward(dlnet,dlimg);Compare the sizes of the spatial dimensions of the layer output with the dimensions of the coefficients at the corresponding decomposition level. Because the reconstruction level is greater than 0 and the difference between the analysis and reconstruction levels is greater than 1, the sizes are equal. The layer effectively sets FullTree to true which removes any ambiguity regarding the size of the reconstruction.
[size(netOutput,[1 2]); size(D{rlevel},[1 2])]ans = 2×2
34 36
34 36
Reconstruction Level is 0
Use the dldwt function to obtain the differentiable DWT of the image down to level 2. Specify the db2 wavelet and zero padding as the extension mode. Set FullTree to true to obtain the full wavelet decomposition.
wv = "db2"; alevel = 2; bdy = "zeropad"; [A,D] = dldwt(dlimg, ... Wavelet=wv, ... Level=alevel, ... FullTree=true, ... PaddingMode=bdy);
Use the dlidwt function to obtain the reconstruction at level 0. Compare the dimensions of the reconstruction with the dimensions of the original signal. Even though FullTree is true, the dimensions are different. Because the reconstruction level is 0, ambiguity exists.
rlevel = 0; xrec = dlidwt(A,D, ... Wavelet=wv, ... Level=rlevel, ... PaddingMode=bdy); [size(dlimg,[1 2]) ; size(xrec,[1 2])]
ans = 2×2
127 135
128 136
Because the reconstruction level is 0, the size of the reconstruction must equal the size of the original signal. You can set ExpectedOutputSize to remove the ambiguity.
xrec2 = dlidwt(A,D, ... Wavelet=wv, ... Level=rlevel, ... PaddingMode=bdy, ... ExpectedOutputSize=size(dlimg,[1 2])); [size(dlimg,[1 2]) ; size(xrec2,[1 2])]
ans = 2×2
127 135
127 135
Similarly, to remove the ambiguity in the wavelet pooling layer, you can specify ExpectedOutputSize. Create a Layer array containing an image input layer and the wavelet pooling layer. Convert the layer array to a dlnetwork object. Run the image through the network.
wpl = waveletPooling2dLayer(Wavelet=wv, ... AnalysisLevel=alevel, ... ReconstructionLevel=rlevel, ... Boundary=bdy, ... ExpectedOutputSize=size(dlimg,[1 2])); layers = [ imageInputLayer([size(dlimg,[1 2]),1]) wpl]; dlnet = dlnetwork(layers); dlnetout = forward(dlnet,dlimg); [size(dlimg,[1 2]) ; size(dlnetout,[1 2])]
ans = 2×2
127 135
127 135
Difference Between Analysis and Reconstruction Levels is 1
When the difference between the analysis and reconstruction levels is 1, the pooling layer behaves as if FullTree is false. Use the dldwt function to obtain the differentiable DWT of the image down to level 2. Specify the db2 wavelet and zero padding as the extension mode. Set FullTree to false.
wv = "db2"; alevel = 2; bdy = "zeropad"; [A,D] = dldwt(dlimg, ... Wavelet=wv, ... Level=alevel, ... FullTree=false, ... PaddingMode=bdy);
If the input coefficients are a tensor, the dlidwt function performs a single-level IDWT. Obtain the size of the reconstruction.
xrec = dlidwt(A,D, ... Wavelet=wv, ... PaddingMode=bdy); size(xrec,[1 2])
ans = 1×2
66 70
Use the wavedec2 function to obtain the bookkeeping matrix of an image whose dimensions equal the size of the spatial dimensions of the dlarray object. Use the same input parameters. Use zero padding boundary extension. Extract from the bookkeeping matrix the sizes of the coefficients at level 2. Because FullTree is false, the sizes extracted from the bookkeeping matrix do not equal the sizes of the reconstruction.
origMode = dwtmode("status","nodisplay"); dwtmode("zpd","nodisplay") [~,s] = wavedec2(img,alevel,wv); dwtmode(origMode,"nodisplay") extractedSize = s(end-2+1,:)
extractedSize = 1×2
65 69
Use the dlidwt function to obtain the reconstruction at level 1, but this time specify ExpectedOutputSize to equal the extracted values.
xrec2 = dlidwt(A,D, ... Wavelet=wv, ... PaddingMode=bdy, ... ExpectedOutputSize=extractedSize); size(xrec2,[1 2])
ans = 1×2
65 69
Create a wavelet pooling layer. Set the layer properties to agree with the dldwt and dlidwt input arguments, including the expected output size. Create a Layer array containing an image input layer and the wavelet pooling layer. Convert the layer array to a dlnetwork object. Run the image through the network. Confirm the sizes of the network output equal ExpectedOutputSize.
rlevel = 1; wpl = waveletPooling2dLayer(Wavelet=wv, ... AnalysisLevel=alevel, ... ReconstructionLevel=rlevel, ... Boundary=bdy, ... ExpectedOutputSize=extractedSize); layers = [ imageInputLayer([size(dlimg,[1 2]),1]) wpl]; dlnet = dlnetwork(layers); dlnetout = forward(dlnet,dlimg); size(dlnetout,[1 2])
ans = 1×2
65 69
More About
A discrete wavelet pooling layer performs downsampling by outputting the wavelet approximation at the specified level.
Williams and Li proposed a wavelet pooling scheme for convolutional-based networks [1]. The layer first
obtains the DWT of the input down to level AnalysisLevel. The layer
then uses the coarse-scale lowpass (scaling) coefficients and the detail coefficients at
decomposition levels AnalysisLevel through
ReconstructionLevel+1ReconstructionLevel. In 1-D wavelet
pooling, the layer pools over the "T” (time) dimension. In 2-D wavelet
pooling, the layer pools over the two "S" (spatial) dimensions. This
diagram illustrates the process for 2-D wavelet pooling:
Wavelet Pooling Scheme
Walter and Garcke were the first to propose an adaptive wavelet pooling approach [2]. In adaptive learnable pooling, the main idea is to multiply the coefficients in each wavelet subband by a weight w and use those weighted subbands to obtain the modified approximation. Because each subband has a different weight, the model can learn which subbands are most relevant to the model’s objective. With the exception of the lowpass weight, Walter and Garcke’s approach is conceptually similar to what is done in traditional wavelet denoising. Their scheme uses gradient descent instead of some other method to determine the threshold. The algorithm for 2-D pooling can be described as follows:
Obtain the wavelet decomposition of the signal at level M.
Multiply the coefficients
LLM,LHM,HLM, andHHMby their respective weightswLL,wLH,wHL, andwHH.Compute the inverse DWT to obtain the modified approximation at level
M-1.
For more information about the 2-D DWT, see wavedec2.
The output of waveletPooling2dLayer is a
dlarray object with format "SSCB". The sizes of the
"S" (spatial) dimensions depend on the sizes of the layer input along
the same dimensions and these layer properties: Wavelet,
Boundary, AnalysisLevel, and
ReconstructionLevel. If you want waveletPooling2dLayer to
verify the output size along the time dimension, set
ExpectedOutputSize.
If ReconstructionLevel is not zero and the difference between
AnalysisLevel and ReconstructionLevel is greater
than 1, by default the layer output dimensions always equal the dimensions you can determine
using dldwt or
wavedec2 (see Determine Expected Output Size of 2-D Discrete Wavelet Pooling Layer).
If ReconstructionLevel is 0 or the difference
between AnalysisLevel and ReconstructionLevel is
exactly 1, there is ambiguity regarding what the layer output dimensions should be. The
dimensions of the coefficients of a single-level forward 2-D DWT is
ceil(sz/2), where sz are the
dimensions of the input, for periodic boundary extension, and
floor((sz+LF-1)/2), where
LF is the filter length, for reflection or zero padding boundary
extension (see wavedec). The floor and ceiling functions
introduce the ambiguity. When performing a single-level inverse 2-D DWT, the layer cannot
determine the dimensions of the original data that was decomposed. In this scenario, you can
use dldwt or
wavedec2 to obtain the expected output
sizes. Otherwise, if you do not set ExpectedOutputSize, the row and
column sizes of the layer output and the sizes you determine might differ by one.
References
[1] Williams, Travis and Robert Y. Li. “Wavelet Pooling for Convolutional Neural Networks.” International Conference on Learning Representations (2018), https://openreview.net/pdf?id=rkhlb8lCZ.
[2] Wolter, Moritz, and Jochen Garcke. “Adaptive Wavelet Pooling for Convolutional Neural Networks.” In Proceedings of The 24th International Conference on Artificial Intelligence and Statistics, edited by Arindam Banerjee and Kenji Fukumizu, vol. 130. PMLR, 2021. https://proceedings.mlr.press/v130/wolter21a.html.
Version History
Introduced in R2026a
See Also
Apps
- Deep Network Designer (Deep Learning Toolbox)
Functions
Objects
waveletPooling1dLayer|modwtLayer|dlarray(Deep Learning Toolbox) |dlnetwork(Deep Learning Toolbox)
Topics
- Practical Introduction to Multiresolution Analysis
- List of Deep Learning Layers (Deep Learning Toolbox)
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)