kfoldLoss
Classification loss for observations not used in training
Description
L = kfoldLoss(CVMdl)CVMdl. That is, for every fold, kfoldLoss estimates
the classification loss for observations that it holds out when it
trains using all other observations.
L contains a classification loss for each
regularization strength in the linear classification models that compose CVMdl.
L = kfoldLoss(CVMdl,Name,Value)Name,Value pair
arguments. For example, indicate which folds to use for the loss calculation
or specify the classification-loss function.
Input Arguments
CVMdl — Cross-validated, binary, linear classification model
ClassificationPartitionedLinear model object
Cross-validated, binary, linear classification model, specified as a ClassificationPartitionedLinear model object. You can create a
ClassificationPartitionedLinear model using fitclinear and specifying any one of the cross-validation, name-value
pair arguments, for example, CrossVal.
To obtain estimates, kfoldLoss applies the same data used to cross-validate the linear
classification model (X and Y).
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Folds — Fold indices to use for classification-score prediction
1:CVMdl.KFold (default) | numeric vector of positive integers
Fold indices to use for classification-score prediction, specified
as the comma-separated pair consisting of 'Folds' and
a numeric vector of positive integers. The elements of Folds must
range from 1 through CVMdl.KFold.
Example: 'Folds',[1 4 10]
Data Types: single | double
LossFun — Loss function
'classiferror' (default) | 'binodeviance' | 'classifcost' | 'exponential' | 'hinge' | 'logit' | 'mincost' | 'quadratic' | function handle
Loss function, specified as the comma-separated pair consisting
of 'LossFun' and a built-in loss function name
or function handle.
The following table lists the available loss functions. Specify one using its corresponding character vector or string scalar.
Value Description "binodeviance"Binomial deviance "classifcost"Observed misclassification cost "classiferror"Misclassified rate in decimal "exponential"Exponential loss "hinge"Hinge loss "logit"Logistic loss "mincost"Minimal expected misclassification cost (for classification scores that are posterior probabilities) "quadratic"Quadratic loss 'mincost'is appropriate for classification scores that are posterior probabilities. For linear classification models, logistic regression learners return posterior probabilities as classification scores by default, but SVM learners do not (seepredict).Specify your own function using function handle notation.
Let
nbe the number of observations inXandKbe the number of distinct classes (numel(Mdl.ClassNames),Mdlis the input model). Your function must have this signaturewhere:lossvalue =lossfun(C,S,W,Cost)The output argument
lossvalueis a scalar.You choose the function name (
lossfun).Cis ann-by-Klogical matrix with rows indicating which class the corresponding observation belongs. The column order corresponds to the class order inMdl.ClassNames.Construct
Cby settingC(p,q) = 1if observationpis in classq, for each row. Set all other elements of rowpto0.Sis ann-by-Knumeric matrix of classification scores. The column order corresponds to the class order inMdl.ClassNames.Sis a matrix of classification scores, similar to the output ofpredict.Wis ann-by-1 numeric vector of observation weights. If you passW, the software normalizes them to sum to1.Costis a K-by-Knumeric matrix of misclassification costs. For example,Cost = ones(K) - eye(K)specifies a cost of0for correct classification, and1for misclassification.
Specify your function using
'LossFun',@.lossfun
Data Types: char | string | function_handle
Mode — Loss aggregation level
'average' (default) | 'individual'
Loss aggregation level, specified as the comma-separated pair
consisting of 'Mode' and 'average' or 'individual'.
| Value | Description |
|---|---|
'average' | Returns losses averaged over all folds |
'individual' | Returns losses for each fold |
Example: 'Mode','individual'
Output Arguments
L — Cross-validated classification losses
numeric scalar | numeric vector | numeric matrix
Cross-validated classification losses, returned
as a numeric scalar, vector, or matrix. The interpretation of L depends
on LossFun.
Let R be the number of regularizations strengths is the
cross-validated models (stored in
numel(CVMdl.Trained{1}.Lambda)) and
F be the number of folds (stored in
CVMdl.KFold).
If
Modeis'average', thenLis a 1-by-Rvector.L(is the average classification loss over all folds of the cross-validated model that uses regularization strengthj)j.Otherwise,
Lis anF-by-Rmatrix.L(is the classification loss for foldi,j)iof the cross-validated model that uses regularization strengthj.
To estimate L,
kfoldLoss uses the data that created
CVMdl (see X and Y).
Examples
Estimate k-Fold Cross-Validation Classification Error
Load the NLP data set.
load nlpdataX is a sparse matrix of predictor data, and Y is a categorical vector of class labels. There are more than two classes in the data.
The models should identify whether the word counts in a web page are from the Statistics and Machine Learning Toolbox™ documentation. So, identify the labels that correspond to the Statistics and Machine Learning Toolbox™ documentation web pages.
Ystats = Y == 'stats';Cross-validate a binary, linear classification model that can identify whether the word counts in a documentation web page are from the Statistics and Machine Learning Toolbox™ documentation.
rng(1); % For reproducibility CVMdl = fitclinear(X,Ystats,'CrossVal','on');
CVMdl is a ClassificationPartitionedLinear model. By default, the software implements 10-fold cross validation. You can alter the number of folds using the 'KFold' name-value pair argument.
Estimate the average of the out-of-fold, classification error rates.
ce = kfoldLoss(CVMdl)
ce = 7.6017e-04
Alternatively, you can obtain the per-fold classification error rates by specifying the name-value pair 'Mode','individual' in kfoldLoss.
Specify Custom Classification Loss
Load the NLP data set. Preprocess the data as in Estimate k-Fold Cross-Validation Classification Error, and transpose the predictor data.
load nlpdata Ystats = Y == 'stats'; X = X';
Cross-validate a binary, linear classification model using 5-fold cross-validation. Optimize the objective function using SpaRSA. Specify that the predictor observations correspond to columns.
rng(1) % For reproducibility CVMdl = fitclinear(X,Ystats,'Solver','sparsa','KFold',5, ... 'ObservationsIn','columns'); CMdl = CVMdl.Trained{1};
CVMdl is a ClassificationPartitionedLinear model. It contains the property Trained, which is a 5-by-1 cell array holding a ClassificationLinear models that the software trained using the training set of each fold.
Create an anonymous function that measures linear loss, that is,
is the weight for observation j, is response j (-1 for the negative class, and 1 otherwise), and is the raw classification score of observation j. Custom loss functions must be written in a particular form. For rules on writing a custom loss function, see the LossFun name-value pair argument. Because the function does not use classification cost, use ~ to have kfoldLoss ignore its position.
linearloss = @(C,S,W,~)sum(-W.*sum(S.*C,2))/sum(W);
Estimate the average cross-validated classification loss using the linear loss function. Also, obtain the loss for each fold.
ce = kfoldLoss(CVMdl,'LossFun',linearloss)ce = -8.0982
ceFold = kfoldLoss(CVMdl,'LossFun',linearloss,'Mode','individual')
ceFold = 5×1
-8.3165
-8.7633
-7.4342
-8.0423
-7.9347
Find Good Lasso Penalty Using k-fold Classification Loss
To determine a good lasso-penalty strength for a linear classification model that uses a logistic regression learner, compare test-sample classification error rates.
Load the NLP data set. Preprocess the data as in Specify Custom Classification Loss.
load nlpdata Ystats = Y == 'stats'; X = X';
Create a set of 11 logarithmically-spaced regularization strengths from through .
Lambda = logspace(-6,-0.5,11);
Cross-validate binary, linear classification models using 5-fold cross-validation, and that use each of the regularization strengths. Optimize the objective function using SpaRSA. Lower the tolerance on the gradient of the objective function to 1e-8.
rng(10); % For reproducibility CVMdl = fitclinear(X,Ystats,'ObservationsIn','columns',... 'KFold',5,'Learner','logistic','Solver','sparsa',... 'Regularization','lasso','Lambda',Lambda,'GradientTolerance',1e-8)
CVMdl =
ClassificationPartitionedLinear
CrossValidatedModel: 'Linear'
ResponseName: 'Y'
NumObservations: 31572
KFold: 5
Partition: [1x1 cvpartition]
ClassNames: [0 1]
ScoreTransform: 'none'
Extract a trained linear classification model.
Mdl1 = CVMdl.Trained{1}Mdl1 =
ClassificationLinear
ResponseName: 'Y'
ClassNames: [0 1]
ScoreTransform: 'logit'
Beta: [34023x11 double]
Bias: [-13.2936 -13.2936 -13.2936 -13.2936 -13.2936 -6.8954 -5.4359 -4.7170 -3.4108 -3.1566 -2.9792]
Lambda: [1.0000e-06 3.5481e-06 1.2589e-05 4.4668e-05 1.5849e-04 5.6234e-04 0.0020 0.0071 0.0251 0.0891 0.3162]
Learner: 'logistic'
Mdl1 is a ClassificationLinear model object. Because Lambda is a sequence of regularization strengths, you can think of Mdl as 11 models, one for each regularization strength in Lambda.
Estimate the cross-validated classification error.
ce = kfoldLoss(CVMdl);
Because there are 11 regularization strengths, ce is a 1-by-11 vector of classification error rates.
Higher values of Lambda lead to predictor variable sparsity, which is a good quality of a classifier. For each regularization strength, train a linear classification model using the entire data set and the same options as when you cross-validated the models. Determine the number of nonzero coefficients per model.
Mdl = fitclinear(X,Ystats,'ObservationsIn','columns',... 'Learner','logistic','Solver','sparsa','Regularization','lasso',... 'Lambda',Lambda,'GradientTolerance',1e-8); numNZCoeff = sum(Mdl.Beta~=0);
In the same figure, plot the cross-validated, classification error rates and frequency of nonzero coefficients for each regularization strength. Plot all variables on the log scale.
figure; [h,hL1,hL2] = plotyy(log10(Lambda),log10(ce),... log10(Lambda),log10(numNZCoeff)); hL1.Marker = 'o'; hL2.Marker = 'o'; ylabel(h(1),'log_{10} classification error') ylabel(h(2),'log_{10} nonzero-coefficient frequency') xlabel('log_{10} Lambda') title('Test-Sample Statistics') hold off
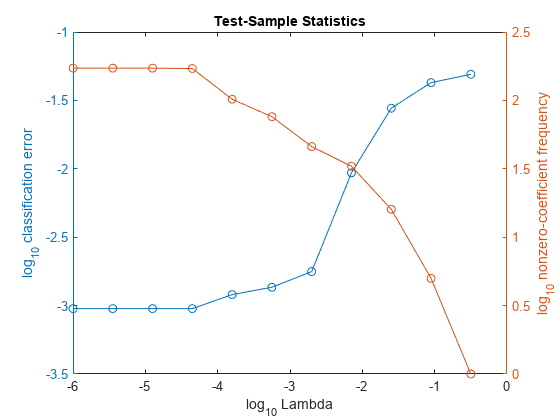
Choose the indexes of the regularization strength that balances predictor variable sparsity and low classification error. In this case, a value between to should suffice.
idxFinal = 7;
Select the model from Mdl with the chosen regularization strength.
MdlFinal = selectModels(Mdl,idxFinal);
MdlFinal is a ClassificationLinear model containing one regularization strength. To estimate labels for new observations, pass MdlFinal and the new data to predict.
More About
Classification Loss
Classification loss functions measure the predictive inaccuracy of classification models. When you compare the same type of loss among many models, a lower loss indicates a better predictive model.
Consider the following scenario.
L is the weighted average classification loss.
n is the sample size.
yj is the observed class label. The software codes it as –1 or 1, indicating the negative or positive class (or the first or second class in the
ClassNamesproperty), respectively.f(Xj) is the positive-class classification score for observation (row) j of the predictor data X.
mj = yjf(Xj) is the classification score for classifying observation j into the class corresponding to yj. Positive values of mj indicate correct classification and do not contribute much to the average loss. Negative values of mj indicate incorrect classification and contribute significantly to the average loss.
The weight for observation j is wj. The software normalizes the observation weights so that they sum to the corresponding prior class probability stored in the
Priorproperty. Therefore,
Given this scenario, the following table describes the supported loss functions that you can specify by using the LossFun name-value argument.
| Loss Function | Value of LossFun | Equation |
|---|---|---|
| Binomial deviance | "binodeviance" | |
| Observed misclassification cost | "classifcost" | where is the class label corresponding to the class with the maximal score, and is the user-specified cost of classifying an observation into class when its true class is yj. |
| Misclassified rate in decimal | "classiferror" | where I{·} is the indicator function. |
| Cross-entropy loss | "crossentropy" |
The weighted cross-entropy loss is where the weights are normalized to sum to n instead of 1. |
| Exponential loss | "exponential" | |
| Hinge loss | "hinge" | |
| Logit loss | "logit" | |
| Minimal expected misclassification cost | "mincost" |
The software computes the weighted minimal expected classification cost using this procedure for observations j = 1,...,n.
The weighted average of the minimal expected misclassification cost loss is |
| Quadratic loss | "quadratic" |
If you use the default cost matrix (whose element value is 0 for correct classification
and 1 for incorrect classification), then the loss values for
"classifcost", "classiferror", and
"mincost" are identical. For a model with a nondefault cost matrix,
the "classifcost" loss is equivalent to the "mincost"
loss most of the time. These losses can be different if prediction into the class with
maximal posterior probability is different from prediction into the class with minimal
expected cost. Note that "mincost" is appropriate only if classification
scores are posterior probabilities.
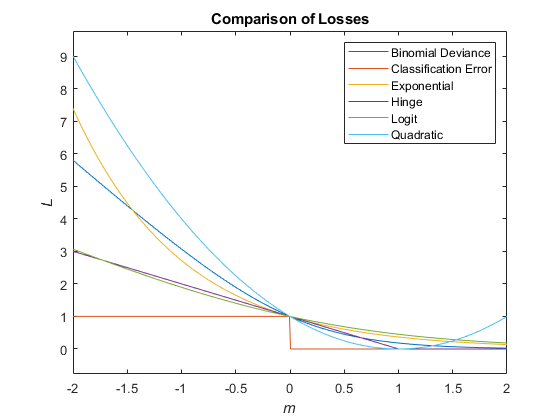
This figure compares the loss functions (except "classifcost",
"crossentropy", and "mincost") over the score
m for one observation. Some functions are normalized to pass through
the point (0,1).
Extended Capabilities
GPU Arrays
Accelerate code by running on a graphics processing unit (GPU) using Parallel Computing Toolbox™.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2016aR2024a: Specify GPU arrays (requires Parallel Computing Toolbox)
kfoldLoss fully supports GPU arrays.
R2023b: Observations with missing predictor values are used in resubstitution and cross-validation computations
Starting in R2023b, the following classification model object functions use observations with missing predictor values as part of resubstitution ("resub") and cross-validation ("kfold") computations for classification edges, losses, margins, and predictions.
In previous releases, the software omitted observations with missing predictor values from the resubstitution and cross-validation computations.
R2022a: kfoldLoss returns a different value for a model with a nondefault cost matrix
If you specify a nondefault cost matrix when you train the input model object, the kfoldLoss function returns a different value compared to previous releases.
The kfoldLoss function uses the
observation weights stored in the W property. Also, the function uses the
cost matrix stored in the Cost property if you specify the
LossFun name-value argument as "classifcost" or
"mincost". The way the function uses the W and
Cost property values has not changed. However, the property values stored in the input model object have changed for a
model with a nondefault cost matrix, so the function might return a different value.
For details about the property value change, see Cost property stores the user-specified cost matrix.
If you want the software to handle the cost matrix, prior
probabilities, and observation weights in the same way as in previous releases, adjust the prior
probabilities and observation weights for the nondefault cost matrix, as described in Adjust Prior Probabilities and Observation Weights for Misclassification Cost Matrix. Then, when you train a
classification model, specify the adjusted prior probabilities and observation weights by using
the Prior and Weights name-value arguments, respectively,
and use the default cost matrix.
See Also
ClassificationPartitionedLinear | ClassificationLinear | kfoldPredict | loss
MATLAB 命令
您点击的链接对应于以下 MATLAB 命令:
请在 MATLAB 命令行窗口中直接输入以执行命令。Web 浏览器不支持 MATLAB 命令。

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)