plsregress
Partial least-squares (PLS) regression
Syntax
Description
[
also returns:XL,YL,XS,YS,BETA,PCTVAR,MSE,stats] = plsregress(X,Y,ncomp)
The predictor scores
XS. Predictor scores are PLS components that are linear combinations of the variables inX.The response scores
YS. Response scores are linear combinations of the responses with which the PLS componentsXShave maximum covariance.The matrix
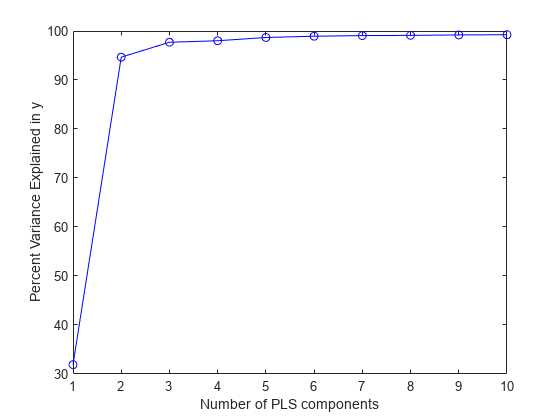
BETAof coefficient estimates for the PLS regression model.The percentage of variance
PCTVARexplained by the regression model.The estimated mean squared errors
MSEfor PLS models withncompcomponents.A structure
statsthat contains the PLS weights, T2 statistic, and predictor and response residuals.
[
specifies options using one or more name-value arguments in addition to any of the
input argument combinations in previous syntaxes. The name-value arguments specify
XL,YL,XS,YS,BETA,PCTVAR,MSE,stats] = plsregress(___,Name,Value)MSE calculation parameters. For example,
'CV',5 calculates the MSE using 5-fold
cross-validation.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Algorithms
plsregress uses the SIMPLS algorithm [1].
If the model fit includes the constant term (intercept), the function first centers
X and Y by subtracting the column means to
get the centered predictor and response variables X0 and
Y0, respectively. However, the function does not rescale the
columns. To perform PLS regression with standardized variables, use zscore to normalize X and Y
(columns of X0 and Y0 are centered to have mean 0
and scaled to have standard deviation 1).
After centering X and Y,
plsregress computes the singular value decomposition (SVD) on
X0'*Y0. The predictor and response loadings
XL and YL are the coefficients obtained
from regressing X0 and Y0 on the predictor score
XS. You can reconstruct the centered data X0
and Y0 using XS*XL' and XS*YL',
respectively.
plsregress initially computes YS as
YS = Y0*YL. By convention [1],
however, plsregress then orthogonalizes each column of
YS with respect to preceding columns of
XS, so that XS'*YS is a lower triangular
matrix.
If the model fit does not include the constant term (intercept),
X and Y are not centered as part of the
fitting process.
References
[1] de Jong, Sijmen. “SIMPLS: An Alternative Approach to Partial Least Squares Regression.” Chemometrics and Intelligent Laboratory Systems 18, no. 3 (March 1993): 251–63. https://doi.org/10.1016/0169-7439(93)85002-X.
[2] Rosipal, Roman, and Nicole Kramer. "Overview and Recent Advances in Partial Least Squares." Subspace, Latent Structure and Feature Selection: Statistical and Optimization Perspectives Workshop (SLSFS 2005), Revised Selected Papers (Lecture Notes in Computer Science 3940). Berlin, Germany: Springer-Verlag, 2006, vol. 3940, pp. 34–51. https://doi.org/10.1007/11752790_2.
[3] Chong, Il-Gyo, and Chi-Hyuck Jun. “Performance of Some Variable Selection Methods When Multicollinearity Is Present.” Chemometrics and Intelligent Laboratory Systems 78, no. 1–2 (July 2005) 103–12. https://doi.org/10.1016/j.chemolab.2004.12.011.
Extended Capabilities
Version History
Introduced in R2008a