Accelerate Signal Feature Extraction and Classification Using a GPU
This example uses signal feature extraction objects to extract multidomain features that can be used to identify faulty bearing signals in mechanical systems. Feature extraction objects enable you to efficiently compute multiple features by reducing the number of times that signals are transformed into a particular domain. The example shows how to compute features and train models using a GPU while running on:
An AMD EPYC 7313P 16-Core Processor @ 3GHz CPU worker
A single NVIDIA® RTX A5000-25.29GB graphical processing unit (GPU)
The acceleration results may vary based on the available hardware resources.
This example extends the workflow described in Machine Learning and Deep Learning Classification Using Signal Feature Extraction Objects. To learn how to extract features and train models in parallel using a parallel pool of workers, see Accelerate Signal Feature Extraction and Classification Using a Parallel Pool of Workers.
Download and Prepare Data
The data set contains acceleration signals collected from rotating machines in a bearing test rig and real-world machines such an oil pump bearing, an intermediate speed bearing, and a planet bearing. There are 34 files in total. The signals in the files are sampled at fs = 48828 Hz. The file names describe the signals they contain:
HealthySignal_*.mat—Healthy signalsInnerRaceFault_*.mat—Signals with inner race faultsOuterRaceFault_*.mat—Signals with outer race faults
Download the data files into a temporary directory. Create a signalDatastore object to access the data in the files and obtain the labels that refer to the signal category.
dataURL = "https://www.mathworks.com/supportfiles/SPT/data/rollingBearingDataset.zip"; datasetFolder = fullfile(tempdir,"rollingBearingDataset"); zipFile = fullfile(tempdir,"rollingBearingDataset.zip"); if ~exist(datasetFolder,"dir") websave(zipFile,dataURL); unzip(zipFile,datasetFolder); end
Create a signalDatastore object to access the data in the files and obtain the labels. Use single-precision arithmetic in the feature extraction and model training steps to reduce memory requirements and computation time.
sds = signalDatastore(datasetFolder,OutputDataType="single");Filenames in the data set includes the labels. Get a list of labels from the filenames in the datastore using the filenames2labels function.
labels = filenames2labels(sds,ExtractBefore=pattern("Signal"|"Fault"));
To accelerate subsequent feature extraction computations using a GPU, create a signalDatastore that returns the variables stored in the files as gpuArray objects. A gpuArray object represents an array stored in GPU memory. Create a datastore that returns gpuArrays with single-precision data.
sdsGPU = signalDatastore(datasetFolder,OutputDataType="single",OutputEnvironment="gpu");
Set Up Feature Extraction Objects
Set up the feature extractors that extract multidomain features from the signals. These features are used to implement machine learning and deep learning solutions to classify signals as healthy, as having inner race faults, or as having outer race faults [1]. Use the signalTimeFeatureExtractor, signalFrequencyFeatureExtractor, and signalTimeFrequencyFeatureExtractor objects to extract features from all the signals.
For time domain, use root-mean-square value, impulse factor, standard deviation, and clearance factor as features.
For frequency domain, use median frequency, band power, power bandwidth, and peak amplitude of the power spectral density (PSD) as features.
For time-frequency domain, use spectral kurtosis [2] of the signal spectrogram as a feature.
Create a signalTimeFeatureExtractor object that extracts the time-domain features using the sample rate fs.
fs = 48828; timeFE = signalTimeFeatureExtractor(SampleRate=fs, ... RMS=true, ... ImpulseFactor=true, ... StandardDeviation=true, ... ClearanceFactor=true);
Create a signalFrequencyFeatureExtractor object that extracts the frequency-domain features.
freqFE = signalFrequencyFeatureExtractor(SampleRate=fs, ... MedianFrequency=true, ... BandPower=true, ... PowerBandwidth=true, ... PeakAmplitude=true);
Create a signalTimeFrequencyFeatureExtractor object that extracts the time-frequency domain features.
timeFreqFE = signalTimeFrequencyFeatureExtractor(SampleRate=fs, ... TimeSpectrum=true); setExtractorParameters(timeFreqFE,"scalogram", ... VoicesPerOctave=16,FrequencyLimits=[50 20000]);
Train SVM Classifier Using Multidomain Features
Extract Multidomain Features
Extract multidomain features using a CPU and a GPU and compare computation time.
Extract features using the CPU and measure the computation time.
tstart = tic;
SVMCPUFeatures = cellfun(@(a,b,c) [real(a) real(b) real(c)],extract(timeFE,sds),extract(freqFE,sds), ...
extract(timeFreqFE,sds),UniformOutput=false);
tCPU = toc(tstart);Accelerate the feature extraction process using a GPU by setting the sdsGPU datastore as input to the extract methods of the feature extractors. Recall that the OutputEnvironment property of this datastore was set to "gpu" for this purpose. Measure the computation time.
device = gpuDevice
device =
CUDADevice with properties:
Name: 'NVIDIA RTX A5000'
Index: 3 (of 4)
ComputeCapability: '8.6'
DriverModel: 'N/A'
TotalMemory: 25294995456 (25.29 GB)
AvailableMemory: 24412668320 (24.41 GB)
DeviceAvailable: true
DeviceSelected: true
Show all properties.
tstart = tic;
SVMGPUFeatures = cellfun(@(a,b,c) [real(a) real(b) real(c)],extract(timeFE,sdsGPU), ...
extract(freqFE,sdsGPU),extract(timeFreqFE,sdsGPU),UniformOutput=false);
wait(device)
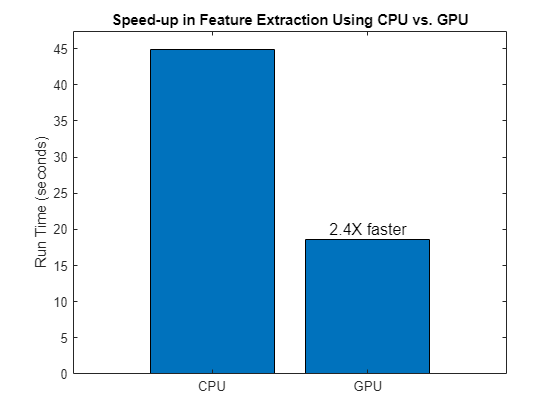
tGPU = toc(tstart);Compare the run times to see the increase in speed you get when you use a GPU for feature extraction.
bar(["CPU" "GPU"],[tCPU tGPU],0.8,FontSize=12, ... Labels=["" num2str(round(tCPU/tGPU,2),2)+ "x faster"]) title("Speed-up in Feature Extraction for SVM Model Using CPU vs. GPU") ylabel("Run Time (seconds)")
Train SVM Classifier Model
Obtain an in-memory feature matrix and use it to train a multiclass SVM classifier. Split the feature matrix into training and testing feature data sets. Obtain their corresponding labels. Reset the random number generator for reproducible results.
featureMatrixCPU = cell2mat(SVMCPUFeatures);
featureMatrixCPU(~isfinite(featureMatrixCPU)) = 0;
rng("default")
cvp = cvpartition(labels,Holdout=0.25);
trainingResponse = labels(cvp.training);
testResponse = labels(cvp.test);Obtain the in-memory feature matrices from training and testing feature data sets. Normalize the in-memory feature matrices and obtain the training and testing feature predictors using the helper function hGetNormalizedSVMFeatureMatrices.
trainMatrixCPU = featureMatrixCPU(cvp.training,:); testMatrixCPU = featureMatrixCPU(cvp.test,:); [trainMatrixCPUNorm,testMatrixCPUNorm] = helperGetNormalizedSVMFeatureMatrices(trainMatrixCPU,testMatrixCPU); trainingPredictorsCPU = array2table(trainMatrixCPUNorm); testPredictors = array2table(testMatrixCPUNorm);
Train a multiclass SVM classifier model using in-memory training feature matrix and its corresponding labels. Obtain the SVM classifier model training time on the CPU.
tStart = tic; [~] = fitcecoc(trainingPredictorsCPU,trainingResponse); tTrainingCPU = toc(tStart);
Accelerate the SVM training process by using GPU features.
featureMatrixGPU = vertcat(SVMGPUFeatures{:});
trainMatrixGPU = featureMatrixGPU(cvp.training,:);
testMatrixGPU = featureMatrixGPU(cvp.test,:);
[trainMatrixGPUNorm,testMatrixGPUNorm] = helperGetNormalizedSVMFeatureMatrices(trainMatrixGPU,testMatrixGPU);
% Create the feature tables
trainingPredictorsGPU = array2table(trainMatrixGPUNorm);
testPredictorsGPU = array2table(testMatrixGPUNorm);Obtain the SVM classifier model training time on the GPU.
tStart = tic; SVMModel = fitcecoc(trainingPredictorsGPU,trainingResponse); wait(device) tTrainingGPU = toc(tStart);
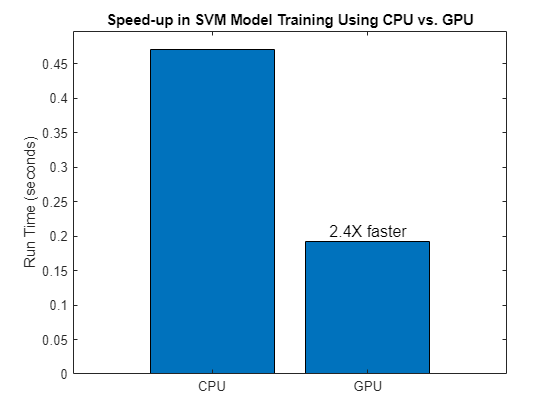
Compare the time it takes to train the model using the CPU and GPU.
bar(["CPU" "GPU"],[tTrainingCPU tTrainingGPU],0.8,FontSize=12, ... Labels=["" num2str(round(tTrainingCPU/tTrainingGPU,2),2)+"x faster"]) title("Speed-up in SVM Model Training Using CPU vs. GPU") ylabel("Run Time (seconds)")
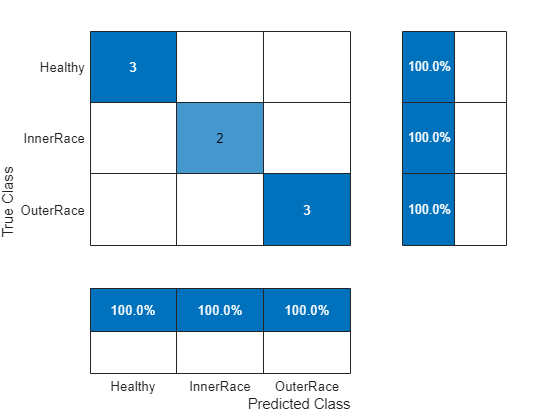
Use the trained SVM classifier and in-memory test features to observe the classifier accuracy.
predictedLabels = predict(SVMModel,testMatrixGPUNorm); figure confusionchart(testResponse,predictedLabels, ... ColumnSummary="column-normalized",RowSummary="row-normalized");
Train LSTM Network Using Features
Set Up Feature Extraction Objects for Training LSTM Network
Each signal in the signalDatastore object sds has around 150,000 samples. Window each signal into 2000-sample signal frames and extract multidomain features from it using all three feature extractors. To window the signals, set the FrameSize for all three feature extractors to 2000.
timeFE.FrameSize = 2000; freqFE.FrameSize = 2000; timeFreqFE.FrameSize = 2000;
Features extracted from frames correspond to a sequence of features over time that have lower dimension than the original signal. The dimension reduction helps the LSTM network to train faster. The workflow follows these steps:
Split the signal datastore and labels into training and test sets.
For each signal in the training and test sets, use all three feature extractor objects to extract features for multiple signal frames. Concatenate the multidomain features to obtain the feature matrix.
Normalize the training and testing feature matrices.
Train the recurrent deep learning network using the labels and feature matrices.
Classify the signals using the trained network.
Split the labels into training and testing sets. Use 70% of the labels for training set and the remaining 30% for testing data. Use splitlabels to partition the labels so that the training and testing data sets hold the same proportion of label values as the entire data set. Obtain the corresponding datastore subsets from the signalDatastore object. Reset the random number generator for reproducible results.
rng("default") splitIndices = splitlabels(labels,0.7,"randomized"); trainIdx = splitIndices{1}; trainLabels = labels(trainIdx);
Extract and Normalize Multidomain Features
Obtain a subset of the files in sds and extract multidomain in-memory training features.
trainDsCPU = subset(sds,trainIdx);
tStart = tic;
trainCPUFeatures = cellfun(@(a,b,c) real([a b c]),extract(timeFE,trainDsCPU), ...
extract(freqFE,trainDsCPU),extract(timeFreqFE,trainDsCPU),UniformOutput=false);
tCPU = toc(tStart);Similarly, obtain a subset of the files in the signalDatastore, sdsGPU, to extract training features using the GPU.
trainDsGPU = subset(sdsGPU,trainIdx);
tStart = tic;
trainGPUFeatures = cellfun(@(a,b,c) real([a b c]),extract(timeFE,trainDsGPU), ...
extract(freqFE,trainDsGPU),extract(timeFreqFE,trainDsGPU),UniformOutput=false);
wait(device)
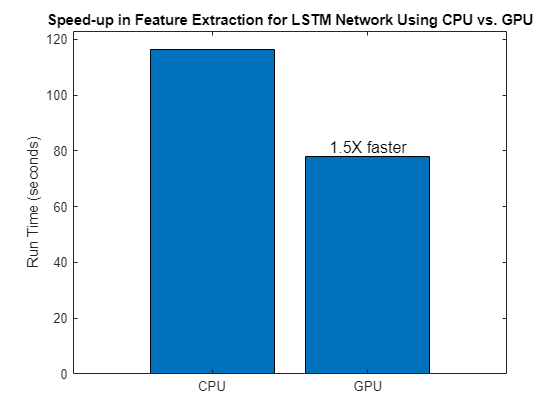
tGPU = toc(tStart);Compare the computation times needed to extract features on the CPU and on the GPU.
bar(["CPU" "GPU"],[tCPU tGPU],0.8,FontSize = 12, ... Labels=["" num2str(round(tCPU/tGPU,2),2)+ "x faster"]) title("Speed-up in Feature Extraction for LSTM Network Using CPU vs. GPU") ylabel("Run Time (seconds)")
Obtain the signalDatastore subsets testDs and testDsGPU for testing from sds and sdsGPU, respectively. Extract the multidomain in-memory and gpuArray test features from testDs and testDsGPU.
testIdx = splitIndices{2};
testLabels = labels(testIdx);
testDs = subset(sds,testIdx);
testCPUFeatures = cellfun(@(a,b,c) [real(a) real(b) real(c)],extract(timeFE,testDs), ...
extract(freqFE,testDs),extract(timeFreqFE,testDs),UniformOutput=false);
testDsGPU = subset(sdsGPU,testIdx);
testGPUFeatures = cellfun(@(a,b,c) [real(a) real(b) real(c)],extract(timeFE,testDsGPU), ...
extract(freqFE,testDsGPU),extract(timeFreqFE,testDsGPU),UniformOutput=false);Normalize the training and testing feature matrices extracted using a CPU and a GPU.
trainCPUFeaturesNorm = ... helperGetNormalizedLSTMFeatureMatrices(trainCPUFeatures); [trainGPUFeaturesNorm,testGPUFeaturesNorm] = ... helperGetNormalizedLSTMFeatureMatrices(trainGPUFeatures,testGPUFeatures);
Train LSTM network
Train an LSTM network using the normalized training features and their corresponding labels.
numFeatures = size(trainCPUFeatures{1},2);
numClasses = 3;
layers = [ ...
sequenceInputLayer(numFeatures)
lstmLayer(50,OutputMode="last")
fullyConnectedLayer(numClasses)
softmaxLayer];
options = trainingOptions("adam", ...
Shuffle="every-epoch", ...
Plots="none", ...
ExecutionEnvironment="cpu", ...
MaxEpochs=100, ...
Verbose=false);
tStart = tic;
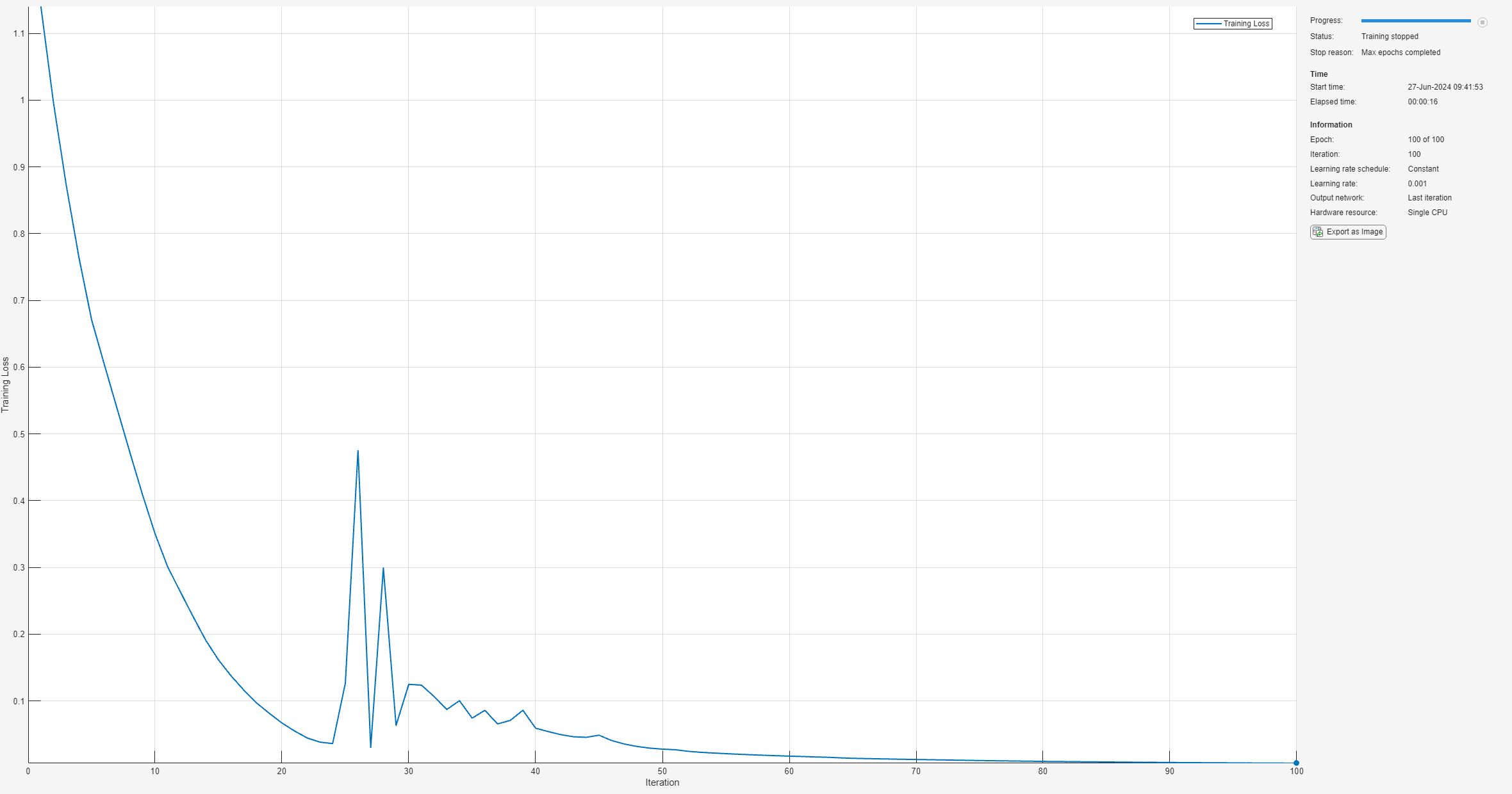
netCPU = trainnet(trainCPUFeaturesNorm,trainLabels,layers,"crossentropy",options);
tTrainingCPU = toc(tStart);Accelerate the training process by setting ExecutionEnvironment to "gpu" in the training options for the network. Compare the time it takes to train the network using the CPU and GPU.
options.ExecutionEnvironment = "gpu";Obtain the LSTM network training time using the GPU features.
tStart = tic;
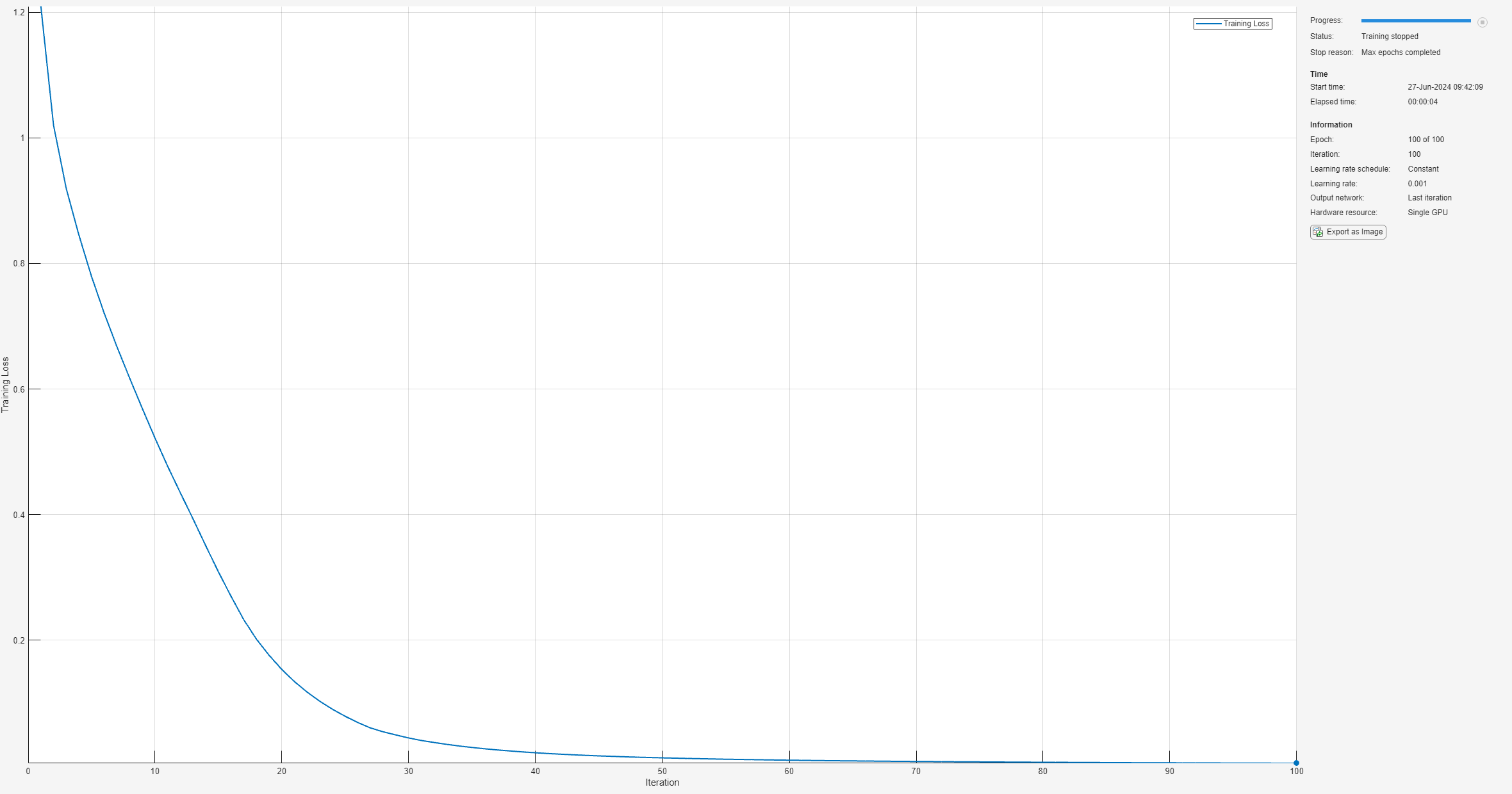
netGPU = trainnet(trainGPUFeaturesNorm,trainLabels,layers,"crossentropy",options);
wait(device)
tTrainingGPU = toc(tStart);Compare the LSTM network training times using in-memory and gpuArray features.
bar(["CPU" "GPU"],[tTrainingCPU tTrainingGPU],0.8,FontSize=12, ... Labels=["" num2str(round(tTrainingCPU/tTrainingGPU,2),2)+"x faster"]) title("Speed-up in LSTM Network Training Using CPU vs. GPU") ylabel("Run Time (seconds)")
Use the trained network to classify the signals in the test data set and analyze the accuracy of the network.
scores = minibatchpredict(netGPU,testGPUFeaturesNorm); classNames = categories(labels); predTest = scores2label(scores,classNames); figure cm = confusionchart(testLabels,predTest, ... ColumnSummary="column-normalized",RowSummary="row-normalized");
References
[1] Caesarendra, Wahyu, and Tegoeh Tjahjowidodo. “A Review of Feature Extraction Methods in Vibration-Based Condition Monitoring and Its Application for Degradation Trend Estimation of Low-Speed Slew Bearing.” Machines 5, no. 4 (December 2017): 21. https://doi.org/10.3390/machines5040021
[2] Tian, Jing, Carlos Morillo, Michael H. Azarian, and Michael Pecht. “Motor Bearing Fault Detection Using Spectral Kurtosis-Based Feature Extraction Coupled With K-Nearest Neighbor Distance Analysis.” IEEE Transactions on Industrial Electronics 63, no. 3 (March 2016): 1793–1803. https://doi.org/10.1109/TIE.2015.2509913
Helper Function
helperGetNormalizedSVMFeatureMatrices – This function normalizes the training and test feature matrices using the mean and standard deviation of the training feature matrix. These normalized features are going to be used for training an SVM model.
function [trainMatrixNorm,testMatrixNorm] = helperGetNormalizedSVMFeatureMatrices(trainMatrix,testMatrix) % Compute normalization parameters from training data featureMean = mean(trainMatrix,1,"omitnan"); featureStd = std(trainMatrix,0,1,"omitnan"); % Normalize both using TRAINING parameters trainMatrixNorm = (trainMatrix-featureMean)./ featureStd; trainMatrixNorm(~isfinite(trainMatrixNorm)) = 0; testMatrixNorm = (testMatrix-featureMean)./ featureStd; testMatrixNorm(~isfinite(testMatrixNorm)) = 0; end
helperGetNormalizedLSTMFeatureMatrices – This function normalizes the training and test feature matrices using the mean and standard deviation of the training feature matrix. The normalized features are going to be used for training an LSTM network.
function [trainFeaturesNorm,testFeaturesNorm] = helperGetNormalizedLSTMFeatureMatrices(trainFeatures,testFeatures) % Compute normalization parameters from training data trainMatrix = cell2mat(trainFeatures); featureMean = mean(trainMatrix,1,"omitnan"); featureStd = std(trainMatrix,0,1,"omitnan"); % Handle zero-variance features zeroVarIdx = featureStd == 0; featureStd(zeroVarIdx) = 1; % Avoid division by zero % Normalize training sequences trainFeaturesNorm = cell(size(trainFeatures)); for i = 1:numel(trainFeatures) trainFeaturesNorm{i} = (trainFeatures{i}-featureMean)./ featureStd; trainFeaturesNorm{i}(~isfinite(trainFeaturesNorm{i})) = 0; end if nargin == 2 % Normalize test sequences using TRAINING parameters testFeaturesNorm = cell(size(testFeatures)); for i = 1:numel(testFeatures) testFeaturesNorm{i} = (testFeatures{i}-featureMean)./ featureStd; testFeaturesNorm{i}(~isfinite(testFeaturesNorm{i})) = 0; end else testFeaturesNorm = []; end end
See Also
Functions
codegen(MATLAB Coder) |confusionchart(Deep Learning Toolbox) |signalDatastore|splitlabels|trainingOptions(Deep Learning Toolbox) |trainnet(Deep Learning Toolbox) |transform