LinearModel
Linear regression model
Description
LinearModel is a fitted linear regression model object. A
regression model describes the relationship between a response and predictors. The
linearity in a linear regression model refers to the linearity of the predictor
coefficients.
Use the properties of a LinearModel object to investigate a fitted
linear regression model. The object properties include information about coefficient
estimates, summary statistics, fitting method, and input data. Use the object functions
to predict responses and to modify, evaluate, and visualize the linear regression
model.
Creation
Create a LinearModel object by using fitlm or stepwiselm.
fitlm fits a linear regression model to
data using a fixed model specification. Use addTerms, removeTerms, or step to add or remove terms from the model. Alternatively, use stepwiselm to fit a model using stepwise linear regression.
Properties
Coefficient Estimates
This property is read-only.
Covariance matrix of coefficient estimates, represented as a
p-by-p matrix of numeric values. p
is the number of coefficients in the fitted model, as given by
NumCoefficients.
For details, see Coefficient Standard Errors and Confidence Intervals.
Data Types: single | double
This property is read-only.
Coefficient names, represented as a cell array of character vectors, each containing the name of the corresponding term.
Data Types: cell
This property is read-only.
Coefficient values, specified as a table.
Coefficients contains one row for each coefficient and these
columns:
Estimate— Estimated coefficient valueSE— Standard error of the estimatetStat— t-statistic for a two-sided test with the null hypothesis that the coefficient is zeropValue— p-value for the t-statistic
Use anova (only for a linear regression model) or
coefTest to perform other tests on the coefficients. Use
coefCI to find the confidence intervals of the coefficient
estimates.
To obtain any of these columns as a vector, index into the property
using dot notation. For example, obtain the estimated coefficient vector in the model
mdl:
beta = mdl.Coefficients.Estimate
Data Types: table
This property is read-only.
Number of model coefficients, represented as a positive integer.
NumCoefficients includes coefficients that are set to zero when
the model terms are rank deficient.
Data Types: double
This property is read-only.
Number of estimated coefficients in the model, specified as a positive integer.
NumEstimatedCoefficients does not include coefficients that are
set to zero when the model terms are rank deficient.
NumEstimatedCoefficients is the degrees of freedom for
regression.
Data Types: double
Summary Statistics
This property is read-only.
Degrees of freedom for the error (residuals), equal to the number of observations minus the number of estimated coefficients, represented as a positive integer.
Data Types: double
This property is read-only.
Observation diagnostics, specified as a table that contains one row for each observation and the columns described in this table.
| Column | Meaning | Description |
|---|---|---|
Leverage | Diagonal elements of HatMatrix | Leverage for each observation indicates to what
extent the fit is determined by the observed predictor values. A value
close to 1 indicates that the fit is largely
determined by that observation, with little contribution from the other
observations. A value close to 0 indicates that the
fit is largely determined by the other observations. For a model with
P coefficients and N
observations, the average value of Leverage is
P/N. A Leverage value greater
than 2*P/N indicates high leverage. |
CooksDistance | Cook's distance | CooksDistance is a measure of scaled change in
fitted values. An observation with CooksDistance
greater than three times the mean Cook's distance can be an outlier. |
Dffits | Delete-1 scaled differences in fitted values | Dffits is the scaled change in the fitted values
for each observation that results from excluding that observation from
the fit. Values greater than 2*sqrt(P/N) in absolute
value can be considered influential. |
S2_i | Delete-1 variance | S2_i is a set of residual variance estimates
obtained by deleting each observation in turn. These estimates can be
compared with the mean squared error (MSE) value, stored in the
MSE property. |
CovRatio | Delete-1 ratio of determinant of covariance | CovRatio is the ratio of the determinant of the
coefficient covariance matrix, with each observation deleted in turn, to
the determinant of the covariance matrix for the full model. Values
greater than 1 + 3*P/N or less than
1 – 3*P/N indicate influential
points. |
Dfbetas | Delete-1 scaled differences in coefficient estimates | Dfbetas is an
N-by-P matrix of the scaled
change in the coefficient estimates that results from excluding each
observation in turn. Values greater than 3/sqrt(N) in
absolute value indicate that the observation has a significant influence
on the corresponding coefficient. |
HatMatrix | Projection matrix to compute fitted from observed
responses | HatMatrix is an
N-by-N matrix such that
Fitted = HatMatrix*Y, where
Y is the response vector and
Fitted is the vector of fitted response values.
|
Diagnostics contains information that is helpful in finding
outliers and influential observations. Delete-1 diagnostics capture the changes that
result from excluding each observation in turn from the fit. For more details, see Hat Matrix and Leverage, Cook’s Distance, and Delete-1 Statistics.
Use plotDiagnostics to plot observation
diagnostics.
Rows not used in the fit because of missing values (in
ObservationInfo.Missing) or excluded values (in
ObservationInfo.Excluded) contain NaN values
in the CooksDistance, Dffits,
S2_i, and CovRatio columns and zeros in the
Leverage, Dfbetas, and
HatMatrix columns.
To obtain any of these columns as an array, index into the property using dot
notation. For example, obtain the delete-1 variance vector in the model
mdl:
S2i = mdl.Diagnostics.S2_i;
Data Types: table
This property is read-only.
Fitted (predicted) response values based on the input data, represented as an
n-by-1 numeric vector. n is the number of
observations in the input data. Use predict to calculate predictions for other predictor values, or to compute
confidence bounds on Fitted.
Data Types: single | double
This property is read-only.
Loglikelihood of response values, specified as a numeric value, based
on the assumption that each response value follows a normal
distribution. The mean of the normal distribution is the fitted
(predicted) response value, and the variance is the
MSE.
Data Types: single | double
This property is read-only.
Criterion for model comparison, represented as a structure with these fields:
AIC— Akaike information criterion.AIC = –2*logL + 2*m, wherelogLis the loglikelihood andmis the number of estimated parameters.AICc— Akaike information criterion corrected for the sample size.AICc = AIC + (2*m*(m + 1))/(n – m – 1), wherenis the number of observations.BIC— Bayesian information criterion.BIC = –2*logL + m*log(n).CAIC— Consistent Akaike information criterion.CAIC = –2*logL + m*(log(n) + 1).
Information criteria are model selection tools that you can use to compare multiple models fit to the same data. These criteria are likelihood-based measures of model fit that include a penalty for complexity (specifically, the number of parameters). Different information criteria are distinguished by the form of the penalty.
When you compare multiple models, the model with the lowest information criterion value is the best-fitting model. The best-fitting model can vary depending on the criterion used for model comparison.
To obtain any of the criterion values as a scalar, index into the property using dot
notation. For example, obtain the AIC value aic in the model
mdl:
aic = mdl.ModelCriterion.AIC
Data Types: struct
This property is read-only.
F-statistic of the regression model, specified as a structure. The
ModelFitVsNullModel structure contains these fields:
Fstats— F-statistic of the fitted model versus the null modelPvalue— p-value for the F-statisticNullModel— null model type
Data Types: struct
This property is read-only.
Mean squared error (residuals), specified as a numeric value.
MSE = SSE / DFE,
where MSE is the mean squared error, SSE is the sum of squared errors, and DFE is the degrees of freedom.
Data Types: single | double
This property is read-only.
Residuals for the fitted model, specified as a table that contains one row for each observation and the columns described in this table.
| Column | Description |
|---|---|
Raw | Observed minus fitted values |
Pearson | Raw residuals divided by the root mean squared error (RMSE) |
Standardized | Raw residuals divided by their estimated standard deviation |
Studentized | Raw residual divided by an independent estimate of the residual standard deviation. The residual for observation i is divided by an estimate of the error standard deviation based on all observations except observation i. |
Use plotResiduals to create a plot of the residuals. For details, see
Residuals.
Rows not used in the fit because of missing values (in
ObservationInfo.Missing) or excluded values (in
ObservationInfo.Excluded) contain NaN values.
To obtain any of these columns as a vector, index into the property using dot notation.
For example, obtain the raw residual vector r in the model
mdl:
r = mdl.Residuals.Raw
Data Types: table
This property is read-only.
Root mean squared error (residuals), specified as a numeric value.
RMSE = sqrt(MSE),
where RMSE is the root mean squared error and MSE is the mean squared error.
Data Types: single | double
This property is read-only.
R-squared value for the model, specified as a structure with two fields:
Ordinary— Ordinary (unadjusted) R-squaredAdjusted— R-squared adjusted for the number of coefficients
The R-squared value is the proportion of the total sum of squares explained by the
model. The ordinary R-squared value relates to the SSR and
SST properties:
Rsquared = SSR/SST,
where SST is the total sum of squares, and
SSR is the regression sum of squares.
For details, see Coefficient of Determination (R-Squared).
To obtain either of these values as a scalar, index into the property using dot
notation. For example, obtain the adjusted R-squared value in the model
mdl:
r2 = mdl.Rsquared.Adjusted
Data Types: struct
This property is read-only.
Sum of squared errors (residuals), specified as a numeric value. If the model was
trained with observation weights, the sum of squares in the SSE
calculation is the weighted sum of squares.
For a linear model with an intercept, the Pythagorean theorem implies
SST = SSE + SSR,
where SST is the total sum of squares,
SSE is the sum of squared errors, and SSR
is the regression sum of squares.
For more information on the calculation of SST for a robust
linear model, see SST.
Data Types: single | double
This property is read-only.
Regression sum of squares, specified as a numeric value.
SSR is equal to the sum of the squared deviations between the fitted
values and the mean of the response. If the model was trained with observation weights, the
sum of squares in the SSR calculation is the weighted sum of
squares.
For a linear model with an intercept, the Pythagorean theorem implies
SST = SSE + SSR,
where SST is the total sum of squares,
SSE is the sum of squared errors, and SSR is the
regression sum of squares.
For more information on the calculation of SST for a robust linear
model, see SST.
Data Types: single | double
This property is read-only.
Total sum of squares, specified as a numeric value. SST is equal
to the sum of squared deviations of the response vector y from the
mean(y). If the model was trained with observation weights, the
sum of squares in the SST calculation is the weighted sum of
squares.
For a linear model with an intercept, the Pythagorean theorem implies
SST = SSE + SSR,
where SST is the total sum of squares,
SSE is the sum of squared errors, and SSR
is the regression sum of squares.
For a robust linear model, SST is not calculated as the sum of
squared deviations of the response vector y from the
mean(y). It is calculated as SST = SSE +
SSR.
Data Types: single | double
Fitting Method
This property is read-only.
Robust fit information, specified as a structure with the fields described in this table.
| Field | Description |
|---|---|
WgtFun | Robust weighting function, such as 'bisquare' (see
'RobustOpts') |
Tune | Tuning constant. This field is empty ([]) if
WgtFun is 'ols' or if
WgtFun is a function handle for a custom weight
function with the default tuning constant 1. |
Weights | Vector of weights used in the final iteration of robust fit. This
field is empty for a CompactLinearModel
object. |
This structure is empty unless you fit the model using robust regression.
Data Types: struct
This property is read-only.
Stepwise fitting information, specified as a structure with the fields described in this table.
| Field | Description |
|---|---|
Start | Formula representing the starting model |
Lower | Formula representing the lower bound model. The terms in
Lower must remain in the model. |
Upper | Formula representing the upper bound model. The model cannot contain
more terms than Upper. |
Criterion | Criterion used for the stepwise algorithm, such as
'sse' |
PEnter | Threshold for Criterion to add a term |
PRemove | Threshold for Criterion to remove a term |
History | Table representing the steps taken in the fit |
The History table contains one row for each step, including the
initial fit, and the columns described in this table.
| Column | Description |
|---|---|
Action | Action taken during the step:
|
TermName |
|
Terms | Model specification in a Terms Matrix |
DF | Regression degrees of freedom after the step |
delDF | Change in regression degrees of freedom from the previous step (negative for steps that remove a term) |
Deviance | Deviance (residual sum of squares) at the step (only for a generalized linear regression model) |
FStat | F-statistic that leads to the step |
PValue | p-value of the F-statistic |
The structure is empty unless you fit the model using stepwise regression.
Data Types: struct
Input Data
This property is read-only after object creation.
Model information, represented as a LinearFormula object.
Display the formula of the fitted model mdl using dot
notation:
mdl.Formula
This property is read-only.
Number of observations the fitting function used in fitting, specified
as a positive integer. NumObservations is the
number of observations supplied in the original table, dataset,
or matrix, minus any excluded rows (set with the
'Exclude' name-value pair
argument) or rows with missing values.
Data Types: double
This property is read-only after object creation.
Number of predictor variables used to fit the model, represented as a positive integer.
Data Types: double
This property is read-only after object creation.
Number of variables in the input data, represented as a positive integer.
NumVariables is the number of variables in the original table, or
the total number of columns in the predictor matrix and response vector.
NumVariables also includes any variables not used to fit the model
as predictors or as the response.
Data Types: double
This property is read-only.
Observation information, specified as an n-by-4 table, where
n is equal to the number of rows of input data.
ObservationInfo contains the columns described in this
table.
| Column | Description |
|---|---|
Weights | Observation weights, specified as a numeric value. The default value
is 1. |
Excluded | Indicator of excluded observations, specified as a logical value. The
value is true if you exclude the observation from the
fit by using the 'Exclude' name-value pair
argument. |
Missing | Indicator of missing observations, specified as a logical value. The
value is true if the observation is missing. |
Subset | Indicator of whether or not the fitting function uses the
observation, specified as a logical value. The value is
true if the observation is not excluded or
missing, meaning the fitting function uses the observation. |
To obtain any of these columns as a vector, index into the property using dot
notation. For example, obtain the weight vector w of the model
mdl:
w = mdl.ObservationInfo.Weights
Data Types: table
This property is read-only after object creation.
Observation names, returned as a cell array of character vectors containing the names of the observations used to fit the model.
If the fit is based on a table containing observation names, this property contains those names.
Otherwise, this property is an empty cell array.
Data Types: cell
This property is read-only after object creation.
Names of predictors used to fit the model, represented as a cell array of character vectors.
Data Types: cell
This property is read-only after object creation.
Response variable name, represented as a character vector.
Data Types: char
This property is read-only after object creation.
Information about the variables contained in Variables,
represented as a table with one row for each variable and the columns described
below.
| Column | Description |
|---|---|
Class | Variable class, specified as a cell array of character vectors, such
as 'double' and
'categorical' |
Range | Variable range, specified as a cell array of vectors
|
InModel | Indicator of which variables are in the fitted model, specified as a
logical vector. The value is true if the model
includes the variable. |
IsCategorical | Indicator of categorical variables, specified as a logical vector.
The value is true if the variable is
categorical. |
VariableInfo also includes any variables not used to fit the model
as predictors or as the response.
Data Types: table
This property is read-only after object creation.
Names of the variables, returned as a cell array of character vectors.
If the fit is based on a table, this property contains the names of the variables in the table.
If the fit is based on a predictor matrix and response vector, this property contains the values specified by the
VarNamesname-value argument of the fitting method. The default value ofVarNamesis{'x1','x2',...,'xn','y'}.
VariableNames also includes any variables not used to fit the model
as predictors or as the response.
Data Types: cell
This property is read-only after object creation.
Input data, returned as a table. Variables contains both
predictor and response values.
If the fit is based on a table, this property contains all the data from the table.
Otherwise, this property is a table created from the input data matrix
Xand the response vectory.
Variables also includes any variables not used to fit the model as
predictors or as the response.
Data Types: table
Object Functions
compact | Compact linear regression model |
addTerms | Add terms to linear regression model |
removeTerms | Remove terms from linear regression model |
step | Improve linear regression model by adding or removing terms |
anova | Analysis of variance for linear regression model |
coefCI | Confidence intervals of coefficient estimates of linear regression model |
coefTest | Linear hypothesis test on linear regression model coefficients |
dwtest | Durbin-Watson test with linear regression model object |
partialDependence | Compute partial dependence |
plot | Scatter plot or added variable plot of linear regression model |
plotAdded | Added variable plot of linear regression model |
plotAdjustedResponse | Adjusted response plot of linear regression model |
plotDiagnostics | Plot observation diagnostics of linear regression model |
plotEffects | Plot main effects of predictors in linear regression model |
plotInteraction | Plot interaction effects of two predictors in linear regression model |
plotPartialDependence | Create partial dependence plot (PDP) and individual conditional expectation (ICE) plots |
plotResiduals | Plot residuals of linear regression model |
plotSlice | Plot of slices through fitted linear regression surface |
gather | Gather properties of Statistics and Machine Learning Toolbox object from GPU |
optimizeResponse | Predictor and response values at response surface maximum or minimum of linear regression model |
matchResponse | Predictor values at specified response value for linear regression model |
Examples
Fit a linear regression model using a matrix input data set.
Load the carsmall data set, a matrix input data set.
load carsmall
X = [Weight,Horsepower,Acceleration];Fit a linear regression model by using fitlm.
mdl = fitlm(X,MPG)
mdl =
Linear regression model:
y ~ 1 + x1 + x2 + x3
Estimated Coefficients:
Estimate SE tStat pValue
__________ _________ _________ __________
(Intercept) 47.977 3.8785 12.37 4.8957e-21
x1 -0.0065416 0.0011274 -5.8023 9.8742e-08
x2 -0.042943 0.024313 -1.7663 0.08078
x3 -0.011583 0.19333 -0.059913 0.95236
Number of observations: 93, Error degrees of freedom: 89
Root Mean Squared Error: 4.09
R-squared: 0.752, Adjusted R-Squared: 0.744
F-statistic vs. constant model: 90, p-value = 7.38e-27
The model display includes the model formula, estimated coefficients, and model summary statistics.
The model formula in the display, y ~ 1 + x1 + x2 + x3, corresponds to . For more information, see Wilkinson Notation.
The model display also shows the estimated coefficient information, which is stored in the Coefficients property. Display the Coefficients property.
mdl.Coefficients
ans=4×4 table
Estimate SE tStat pValue
__________ _________ _________ __________
(Intercept) 47.977 3.8785 12.37 4.8957e-21
x1 -0.0065416 0.0011274 -5.8023 9.8742e-08
x2 -0.042943 0.024313 -1.7663 0.08078
x3 -0.011583 0.19333 -0.059913 0.95236
The Coefficient property includes these columns:
Estimate— Coefficient estimates for each corresponding term in the model. For example, the estimate for the constant term (intercept) is 47.977.SE— Standard error of the coefficients.tStat— t-statistic for each coefficient to test the null hypothesis that the corresponding coefficient is zero against the alternative that it is different from zero, given the other predictors in the model. Note thattStat = Estimate/SE. For example, the t-statistic for the intercept is 47.977/3.8785 = 12.37.pValue— p-value for the t-statistic of the two-sided hypothesis test. For example, the p-value of the t-statistic forx2is greater than 0.05, so this term is not significant at the 5% significance level given the other terms in the model.
The summary statistics of the model are:
Number of observations— Number of rows without anyNaNvalues. For example,Number of observationsis 93 because theMPGdata vector has sixNaNvalues and theHorsepowerdata vector has oneNaNvalue for a different observation, where the number of rows inXandMPGis 100.Error degrees of freedom— n – p, where n is the number of observations, and p is the number of coefficients in the model, including the intercept. For example, the model has four predictors, so theError degrees of freedomis 93 – 4 = 89.Root mean squared error— Square root of the mean squared error, which estimates the standard deviation of the error distribution.R-squaredandAdjusted R-squared— Coefficient of determination and adjusted coefficient of determination, respectively. For example, theR-squaredvalue suggests that the model explains approximately 75% of the variability in the response variableMPG.F-statistic vs. constant model— Test statistic for the F-test on the regression model, which tests whether the model fits significantly better than a degenerate model consisting of only a constant term.p-value— p-value for the F-test on the model. For example, the model is significant with a p-value of 7.3816e-27.
You can find these statistics in the model properties (NumObservations, DFE, RMSE, and Rsquared) and by using the anova function.
anova(mdl,'summary')ans=3×5 table
SumSq DF MeanSq F pValue
______ __ ______ ______ __________
Total 6004.8 92 65.269
Model 4516 3 1505.3 89.987 7.3816e-27
Residual 1488.8 89 16.728
Use plot to create an added variable plot (partial regression leverage plot) for the whole model except the constant (intercept) term.
plot(mdl)
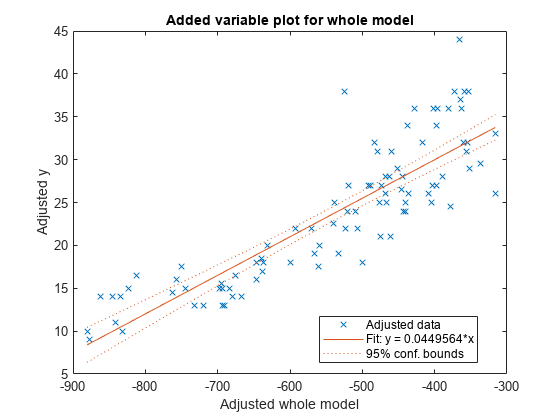
Fit a linear regression model that contains a categorical predictor. Reorder the categories of the categorical predictor to control the reference level in the model. Then, use anova to test the significance of the categorical variable.
Model with Categorical Predictor
Load the carsmall data set and create a linear regression model of MPG as a function of Model_Year. To treat the numeric vector Model_Year as a categorical variable, identify the predictor using the 'CategoricalVars' name-value pair argument.
load carsmall mdl = fitlm(Model_Year,MPG,'CategoricalVars',1,'VarNames',{'Model_Year','MPG'})
mdl =
Linear regression model:
MPG ~ 1 + Model_Year
Estimated Coefficients:
Estimate SE tStat pValue
________ ______ ______ __________
(Intercept) 17.69 1.0328 17.127 3.2371e-30
Model_Year_76 3.8839 1.4059 2.7625 0.0069402
Model_Year_82 14.02 1.4369 9.7571 8.2164e-16
Number of observations: 94, Error degrees of freedom: 91
Root Mean Squared Error: 5.56
R-squared: 0.531, Adjusted R-Squared: 0.521
F-statistic vs. constant model: 51.6, p-value = 1.07e-15
The model formula in the display, MPG ~ 1 + Model_Year, corresponds to
,
where and are indicator variables whose value is one if the value of Model_Year is 76 and 82, respectively. For more information about the model formula, see Wilkinson Notation. The Model_Year variable includes three distinct values, which you can check by using the unique function.
unique(Model_Year)
ans = 3×1
70
76
82
fitlm chooses the smallest value in Model_Year as a reference level ('70') and creates two indicator variables and . The model includes only two indicator variables because the design matrix becomes rank deficient if the model includes three indicator variables (one for each level) and an intercept term.
Model with Full Indicator Variables
You can interpret the model formula of mdl as a model that has three indicator variables without an intercept term:
.
Alternatively, you can create a model that has three indicator variables without an intercept term by manually creating indicator variables and specifying the model formula.
temp_Year = dummyvar(categorical(Model_Year));
Model_Year_70 = temp_Year(:,1);
Model_Year_76 = temp_Year(:,2);
Model_Year_82 = temp_Year(:,3);
tbl = table(Model_Year_70,Model_Year_76,Model_Year_82,MPG);
mdl = fitlm(tbl,'MPG ~ Model_Year_70 + Model_Year_76 + Model_Year_82 - 1')mdl =
Linear regression model:
MPG ~ Model_Year_70 + Model_Year_76 + Model_Year_82
Estimated Coefficients:
Estimate SE tStat pValue
________ _______ ______ __________
Model_Year_70 17.69 1.0328 17.127 3.2371e-30
Model_Year_76 21.574 0.95387 22.617 4.0156e-39
Model_Year_82 31.71 0.99896 31.743 5.2234e-51
Number of observations: 94, Error degrees of freedom: 91
Root Mean Squared Error: 5.56
Choose Reference Level in Model
You can choose a reference level by modifying the order of categories in a categorical variable. First, create a categorical variable Year.
Year = categorical(Model_Year);
Check the order of categories by using the categories function.
categories(Year)
ans = 3×1 cell
{'70'}
{'76'}
{'82'}
If you use Year as a predictor variable, then fitlm chooses the first category '70' as a reference level. Reorder Year by using the reordercats function.
Year_reordered = reordercats(Year,{'76','70','82'});
categories(Year_reordered)ans = 3×1 cell
{'76'}
{'70'}
{'82'}
The first category of Year_reordered is '76'. Create a linear regression model of MPG as a function of Year_reordered.
mdl2 = fitlm(Year_reordered,MPG,'VarNames',{'Model_Year','MPG'})
mdl2 =
Linear regression model:
MPG ~ 1 + Model_Year
Estimated Coefficients:
Estimate SE tStat pValue
________ _______ _______ __________
(Intercept) 21.574 0.95387 22.617 4.0156e-39
Model_Year_70 -3.8839 1.4059 -2.7625 0.0069402
Model_Year_82 10.136 1.3812 7.3385 8.7634e-11
Number of observations: 94, Error degrees of freedom: 91
Root Mean Squared Error: 5.56
R-squared: 0.531, Adjusted R-Squared: 0.521
F-statistic vs. constant model: 51.6, p-value = 1.07e-15
mdl2 uses '76' as a reference level and includes two indicator variables and .
Evaluate Categorical Predictor
The model display of mdl2 includes a p-value of each term to test whether or not the corresponding coefficient is equal to zero. Each p-value examines each indicator variable. To examine the categorical variable Model_Year as a group of indicator variables, use anova. Use the 'components'(default) option to return a component ANOVA table that includes ANOVA statistics for each variable in the model except the constant term.
anova(mdl2,'components')ans=2×5 table
SumSq DF MeanSq F pValue
______ __ ______ _____ __________
Model_Year 3190.1 2 1595.1 51.56 1.0694e-15
Error 2815.2 91 30.936
The component ANOVA table includes the p-value of the Model_Year variable, which is smaller than the p-values of the indicator variables.
Load the hald data set, which measures the effect of cement composition on its hardening heat.
load haldThis data set includes the variables ingredients and heat. The matrix ingredients contains the percent composition of four chemicals present in the cement. The vector heat contains the values for the heat hardening after 180 days for each cement sample.
Fit a robust linear regression model to the data.
mdl = fitlm(ingredients,heat,'RobustOpts','on')
mdl =
Linear regression model (robust fit):
y ~ 1 + x1 + x2 + x3 + x4
Estimated Coefficients:
Estimate SE tStat pValue
________ _______ ________ ________
(Intercept) 60.09 75.818 0.79256 0.4509
x1 1.5753 0.80585 1.9548 0.086346
x2 0.5322 0.78315 0.67957 0.51596
x3 0.13346 0.8166 0.16343 0.87424
x4 -0.12052 0.7672 -0.15709 0.87906
Number of observations: 13, Error degrees of freedom: 8
Root Mean Squared Error: 2.65
R-squared: 0.979, Adjusted R-Squared: 0.969
F-statistic vs. constant model: 94.6, p-value = 9.03e-07
For more details, see the topic Reduce Outlier Effects Using Robust Regression, which compares the results of a robust fit to a standard least-squares fit.
Load the hald data set, which measures the effect of cement composition on its hardening heat.
load haldThis data set includes the variables ingredients and heat. The matrix ingredients contains the percent composition of four chemicals present in the cement. The vector heat contains the values for the heat hardening after 180 days for each cement sample.
Fit a stepwise linear regression model to the data. Specify 0.06 as the threshold for the criterion to add a term to the model.
mdl = stepwiselm(ingredients,heat,'PEnter',0.06)1. Adding x4, FStat = 22.7985, pValue = 0.000576232 2. Adding x1, FStat = 108.2239, pValue = 1.105281e-06 3. Adding x2, FStat = 5.0259, pValue = 0.051687 4. Removing x4, FStat = 1.8633, pValue = 0.2054
mdl =
Linear regression model:
y ~ 1 + x1 + x2
Estimated Coefficients:
Estimate SE tStat pValue
________ ________ ______ __________
(Intercept) 52.577 2.2862 22.998 5.4566e-10
x1 1.4683 0.1213 12.105 2.6922e-07
x2 0.66225 0.045855 14.442 5.029e-08
Number of observations: 13, Error degrees of freedom: 10
Root Mean Squared Error: 2.41
R-squared: 0.979, Adjusted R-Squared: 0.974
F-statistic vs. constant model: 230, p-value = 4.41e-09
By default, the initial model is a constant model. stepwiselm performs forward selection and adds the x4, x1, and x2 terms (in that order), because the corresponding p-values are less than the PEnter value of 0.06. stepwiselm then uses backward elimination and removes x4 from the model because, once x2 is in the model, the p-value of x4 is greater than the default value of PRemove, 0.1.
More About
A terms matrix
T is a t-by-(p + 1) matrix that
specifies the terms in a model, where t is the number of terms,
p is the number of predictor variables, and +1 accounts for the
response variable. The value of T(i,j) is the exponent of variable
j in term i.
For example, suppose that an input includes three predictor variables, x1,
x2, and x3, and the response variable
y in the order x1, x2,
x3, and y. Each row of T
represents one term:
[0 0 0 0]— Constant term (intercept)[0 1 0 0]—x2; equivalently,x1^0 * x2^1 * x3^0[1 0 1 0]—x1*x3[2 0 0 0]—x1^2[0 1 2 0]—x2*(x3^2)
The 0 at the end of each term represents the response variable. In
general, a column vector of zeros in a terms matrix represents the position of the response
variable. If the predictor and response variables are in a matrix and column vector,
respectively, then you must include 0 for the response variable in the
last column of each row.
Alternative Functionality
For reduced computation time on high-dimensional data sets, fit a linear regression model using the
fitrlinearfunction.To regularize a regression, use
fitrlinear,lasso,ridge, orplsregress.fitrlinearregularizes a regression for high-dimensional data sets using lasso or ridge regression.lassoremoves redundant predictors in linear regression using lasso or elastic net.ridgeregularizes a regression with correlated terms using ridge regression.plsregressregularizes a regression with correlated terms using partial least squares.
Extended Capabilities
Usage notes and limitations:
For more information, see Introduction to Code Generation for Statistics and Machine Learning Functions.
Usage notes and limitations:
The object functions of a
LinearModelmodel fully support GPU arrays.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2012a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)