fitcecoc
Fit multiclass models for support vector machines or other classifiers
Syntax
Description
Mdl = fitcecoc(Tbl,ResponseVarName)Tbl and
the class labels in Tbl.ResponseVarName. fitcecoc uses K(K –
1)/2 binary support vector machine (SVM) models using the one-versus-one coding design, where K is
the number of unique class labels (levels). Mdl is
a ClassificationECOC model.
Mdl = fitcecoc(___,Name,Value)Name,Value pair
arguments, using any of the previous syntaxes.
For example, specify different binary learners, a different
coding design, or to cross-validate. It is good practice to cross-validate
using the Kfold Name,Value pair
argument. The cross-validation results determine how well the model
generalizes.
[ also returns
hyperparameter optimization results when you specify
Mdl,HyperparameterOptimizationResults]
= fitcecoc(___,Name,Value)OptimizeHyperparameters and either of the following
conditions apply:
You specify
Learners="linear"orLearners="kernel".HyperparameterOptimizationResultsis aSupervisedLearningBayesianOptimizationobject.You specify
HyperparameterOptimizationOptionsand set theConstraintTypeandConstraintBoundsoptions.HyperparameterOptimizationResultsis anAggregateBayesianOptimizationobject. You can choose to optimize on compact model size or cross-validation loss, and to perform a set of multiple optimization problems that have the same options but different constraint bounds.
When you specify OptimizeHyperparameters
and neither of these conditions apply, the output
HyperparameterOptimizationResults is
[] and, instead, the
HyperparameterOptimizationResults property of
Mdl contains the results.
Note
For a list of supported syntaxes when the input variables are tall arrays, see Tall Arrays.
Examples
Train a multiclass error-correcting output codes (ECOC) model using support vector machine (SVM) binary learners.
Load Fisher's iris data set. Specify the predictor data X and the response data Y.
load fisheriris
X = meas;
Y = species;Train a multiclass ECOC model using the default options.
Mdl = fitcecoc(X,Y)
Mdl =
ClassificationECOC
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
BinaryLearners: {3×1 cell}
CodingName: 'onevsone'
Properties, Methods
Mdl is a ClassificationECOC model. By default, fitcecoc uses SVM binary learners and a one-versus-one coding design. You can access Mdl properties using dot notation.
Display the class names and the coding design matrix.
Mdl.ClassNames
ans = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
CodingMat = Mdl.CodingMatrix
CodingMat = 3×3
1 1 0
-1 0 1
0 -1 -1
A one-versus-one coding design for three classes yields three binary learners. The columns of CodingMat correspond to the learners, and the rows correspond to the classes. The class order is the same as the order in Mdl.ClassNames. For example, CodingMat(:,1) is [1; –1; 0] and indicates that the software trains the first SVM binary learner using all observations classified as 'setosa' and 'versicolor'. Because 'setosa' corresponds to 1, it is the positive class; 'versicolor' corresponds to –1, so it is the negative class.
You can access each binary learner using cell indexing and dot notation.
Mdl.BinaryLearners{1} % The first binary learnerans =
CompactClassificationSVM
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: [-1 1]
ScoreTransform: 'none'
Beta: [4×1 double]
Bias: 1.4492
KernelParameters: [1×1 struct]
Properties, Methods
Compute the resubstitution classification error.
error = resubLoss(Mdl)
error = 0.0067
The classification error on the training data is small, but the classifier might be an overfitted model. You can cross-validate the classifier using crossval and compute the cross-validation classification error instead.
Create a default linear learner template, and then use it to train an ECOC model containing multiple binary linear classification models.
Load the NLP data set.
load nlpdataX is a sparse matrix of predictor data, and Y is a categorical vector of class labels. The data contains 13 classes.
Create a default linear learner template.
t = templateLinear
t =
Fit template for Linear.
Learner: 'svm'
t is a template object for a linear learner. All of the properties of t are empty. When you pass t to a training function, such as fitcecoc for ECOC multiclass classification, the software sets the empty properties to their respective default values. For example, the software sets Type to "classification". To modify the default values see the name-value arguments for templateLinear.
Train an ECOC model consisting of multiple binary linear classification models that identify the software product given the frequency distribution of words on a documentation web page. For faster training time, transpose the predictor data, and specify that observations correspond to columns.
X = X'; rng(1); % For reproducibility Mdl = fitcecoc(X,Y,'Learners',t,'ObservationsIn','columns')
Mdl =
CompactClassificationECOC
ResponseName: 'Y'
ClassNames: [comm dsp ecoder fixedpoint hdlcoder phased physmod simulink stats supportpkg symbolic vision xpc]
ScoreTransform: 'none'
BinaryLearners: {78×1 cell}
CodingMatrix: [13×78 double]
Properties, Methods
Alternatively, you can train an ECOC model containing default linear classification models by specifying "Learners","Linear".
To conserve memory, fitcecoc returns trained ECOC models containing linear classification learners in CompactClassificationECOC model objects.
Cross-validate an ECOC classifier with SVM binary learners, and estimate the generalized classification error.
Load Fisher's iris data set. Specify the predictor data X and the response data Y.
load fisheriris X = meas; Y = species; rng(1); % For reproducibility
Create an SVM template, and standardize the predictors.
t = templateSVM('Standardize',true)t =
Fit template for SVM.
Standardize: 1
t is an SVM template. Most of the template object properties are empty. When training the ECOC classifier, the software sets the applicable properties to their default values.
Train the ECOC classifier, and specify the class order.
Mdl = fitcecoc(X,Y,'Learners',t,... 'ClassNames',{'setosa','versicolor','virginica'});
Mdl is a ClassificationECOC classifier. You can access its properties using dot notation.
Cross-validate Mdl using 10-fold cross-validation.
CVMdl = crossval(Mdl);
CVMdl is a ClassificationPartitionedECOC cross-validated ECOC classifier.
Estimate the generalized classification error.
genError = kfoldLoss(CVMdl)
genError = 0.0400
The generalized classification error is 4%, which indicates that the ECOC classifier generalizes fairly well.
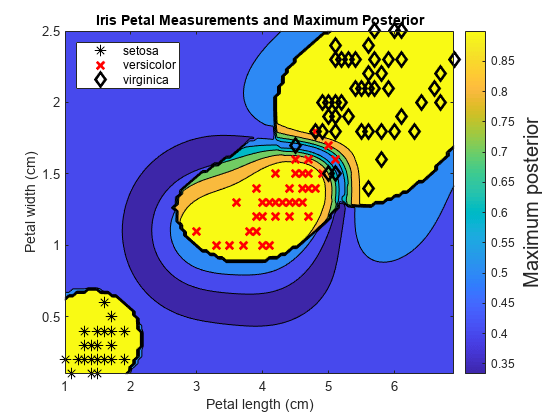
Train an ECOC classifier using SVM binary learners. First predict the training-sample labels and class posterior probabilities. Then predict the maximum class posterior probability at each point in a grid. Visualize the results.
Load Fisher's iris data set. Specify the petal dimensions as the predictors and the species names as the response.
load fisheriris X = meas(:,3:4); Y = species; rng(1); % For reproducibility
Create an SVM template. Standardize the predictors, and specify the Gaussian kernel.
t = templateSVM('Standardize',true,'KernelFunction','gaussian');
t is an SVM template. Most of its properties are empty. When the software trains the ECOC classifier, it sets the applicable properties to their default values.
Train the ECOC classifier using the SVM template. Transform classification scores to class posterior probabilities (which are returned by predict or resubPredict) using the 'FitPosterior' name-value pair argument. Specify the class order using the 'ClassNames' name-value pair argument. Display diagnostic messages during training by using the 'Verbose' name-value pair argument.
Mdl = fitcecoc(X,Y,'Learners',t,'FitPosterior',true,... 'ClassNames',{'setosa','versicolor','virginica'},... 'Verbose',2);
Training binary learner 1 (SVM) out of 3 with 50 negative and 50 positive observations. Negative class indices: 2 Positive class indices: 1 Fitting posterior probabilities for learner 1 (SVM). Training binary learner 2 (SVM) out of 3 with 50 negative and 50 positive observations. Negative class indices: 3 Positive class indices: 1 Fitting posterior probabilities for learner 2 (SVM). Training binary learner 3 (SVM) out of 3 with 50 negative and 50 positive observations. Negative class indices: 3 Positive class indices: 2 Fitting posterior probabilities for learner 3 (SVM).
Mdl is a ClassificationECOC model. The same SVM template applies to each binary learner, but you can adjust options for each binary learner by passing in a cell vector of templates.
Predict the training-sample labels and class posterior probabilities. Display diagnostic messages during the computation of labels and class posterior probabilities by using the 'Verbose' name-value pair argument.
[label,~,~,Posterior] = resubPredict(Mdl,'Verbose',1);Predictions from all learners have been computed. Loss for all observations has been computed. Computing posterior probabilities...
Mdl.BinaryLoss
ans = 'quadratic'
The software assigns an observation to the class that yields the smallest average binary loss. Because all binary learners are computing posterior probabilities, the binary loss function is quadratic.
Display a random set of results.
idx = randsample(size(X,1),10,1); Mdl.ClassNames
ans = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
table(Y(idx),label(idx),Posterior(idx,:),... 'VariableNames',{'TrueLabel','PredLabel','Posterior'})
ans=10×3 table
TrueLabel PredLabel Posterior
______________ ______________ ______________________________________
{'virginica' } {'virginica' } 0.0039319 0.0039866 0.99208
{'virginica' } {'virginica' } 0.017066 0.018262 0.96467
{'virginica' } {'virginica' } 0.014947 0.015855 0.9692
{'versicolor'} {'versicolor'} 2.2197e-14 0.87318 0.12682
{'setosa' } {'setosa' } 0.999 0.00025091 0.00074639
{'versicolor'} {'virginica' } 2.2195e-14 0.059427 0.94057
{'versicolor'} {'versicolor'} 2.2194e-14 0.97002 0.029984
{'setosa' } {'setosa' } 0.999 0.0002499 0.00074741
{'versicolor'} {'versicolor'} 0.0085638 0.98259 0.0088482
{'setosa' } {'setosa' } 0.999 0.00025013 0.00074718
The columns of Posterior correspond to the class order of Mdl.ClassNames.
Define a grid of values in the observed predictor space. Predict the posterior probabilities for each instance in the grid.
xMax = max(X); xMin = min(X); x1Pts = linspace(xMin(1),xMax(1)); x2Pts = linspace(xMin(2),xMax(2)); [x1Grid,x2Grid] = meshgrid(x1Pts,x2Pts); [~,~,~,PosteriorRegion] = predict(Mdl,[x1Grid(:),x2Grid(:)]);
For each coordinate on the grid, plot the maximum class posterior probability among all classes.
contourf(x1Grid,x2Grid,... reshape(max(PosteriorRegion,[],2),size(x1Grid,1),size(x1Grid,2))); h = colorbar; h.YLabel.String = 'Maximum posterior'; h.YLabel.FontSize = 15; hold on gh = gscatter(X(:,1),X(:,2),Y,'krk','*xd',8); gh(2).LineWidth = 2; gh(3).LineWidth = 2; title('Iris Petal Measurements and Maximum Posterior') xlabel('Petal length (cm)') ylabel('Petal width (cm)') axis tight legend(gh,'Location','NorthWest') hold off
Train a one-versus-all ECOC classifier using a GentleBoost ensemble of decision trees with surrogate splits. To speed up training, bin numeric predictors and use parallel computing. Binning is valid only when fitcecoc uses a tree learner. After training, estimate the classification error using 10-fold cross-validation. Note that parallel computing requires Parallel Computing Toolbox™.
Load Sample Data
Load and inspect the arrhythmia data set.
load arrhythmia
[n,p] = size(X)n = 452
p = 279
isLabels = unique(Y); nLabels = numel(isLabels)
nLabels = 13
tabulate(categorical(Y))
Value Count Percent
1 245 54.20%
2 44 9.73%
3 15 3.32%
4 15 3.32%
5 13 2.88%
6 25 5.53%
7 3 0.66%
8 2 0.44%
9 9 1.99%
10 50 11.06%
14 4 0.88%
15 5 1.11%
16 22 4.87%
The data set contains 279 predictors, and the sample size of 452 is relatively small. Of the 16 distinct labels, only 13 are represented in the response (Y). Each label describes various degrees of arrhythmia, and 54.20% of the observations are in class 1.
Train One-Versus-All ECOC Classifier
Create an ensemble template. You must specify at least three arguments: a method, a number of learners, and the type of learner. For this example, specify 'GentleBoost' for the method, 100 for the number of learners, and a decision tree template that uses surrogate splits because there are missing observations.
tTree = templateTree('surrogate','on'); tEnsemble = templateEnsemble('GentleBoost',100,tTree);
tEnsemble is a template object. Most of its properties are empty, but the software fills them with their default values during training.
Train a one-versus-all ECOC classifier using the ensembles of decision trees as binary learners. To speed up training, use binning and parallel computing.
Binning (
'NumBins',50) — When you have a large training data set, you can speed up training (a potential decrease in accuracy) by using the'NumBins'name-value pair argument. This argument is valid only whenfitcecocuses a tree learner. If you specify the'NumBins'value, then the software bins every numeric predictor into a specified number of equiprobable bins, and then grows trees on the bin indices instead of the original data. You can try'NumBins',50first, and then change the'NumBins'value depending on the accuracy and training speed.Parallel computing (
'Options',statset('UseParallel',true)) — With a Parallel Computing Toolbox license, you can speed up the computation by using parallel computing, which sends each binary learner to a worker in the pool. The number of workers depends on your system configuration. When you use decision trees for binary learners,fitcecocparallelizes training using Intel® Threading Building Blocks (TBB) for dual-core systems and above. Therefore, specifying the'UseParallel'option is not helpful on a single computer. Use this option on a cluster.
Additionally, specify that the prior probabilities are 1/K, where K = 13 is the number of distinct classes.
options = statset('UseParallel',true); Mdl = fitcecoc(X,Y,'Coding','onevsall','Learners',tEnsemble,... 'Prior','uniform','NumBins',50,'Options',options);
Starting parallel pool (parpool) using the 'local' profile ... Connected to the parallel pool (number of workers: 6).
Mdl is a ClassificationECOC model.
Cross-Validation
Cross-validate the ECOC classifier using 10-fold cross-validation.
CVMdl = crossval(Mdl,'Options',options);Warning: One or more folds do not contain points from all the groups.
CVMdl is a ClassificationPartitionedECOC model. The warning indicates that some classes are not represented while the software trains at least one fold. Therefore, those folds cannot predict labels for the missing classes. You can inspect the results of a fold using cell indexing and dot notation. For example, access the results of the first fold by entering CVMdl.Trained{1}.
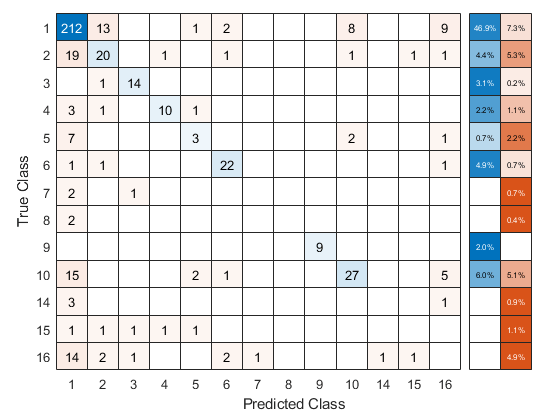
Use the cross-validated ECOC classifier to predict validation-fold labels. You can compute the confusion matrix by using confusionchart. Move and resize the chart by changing the inner position property to ensure that the percentages appear in the row summary.
oofLabel = kfoldPredict(CVMdl,'Options',options); ConfMat = confusionchart(Y,oofLabel,'RowSummary','total-normalized'); ConfMat.InnerPosition = [0.10 0.12 0.85 0.85];
Reproduce Binned Data
Reproduce binned predictor data by using the BinEdges property of the trained model and the discretize function.
X = Mdl.X; % Predictor data Xbinned = zeros(size(X)); edges = Mdl.BinEdges; % Find indices of binned predictors. idxNumeric = find(~cellfun(@isempty,edges)); if iscolumn(idxNumeric) idxNumeric = idxNumeric'; end for j = idxNumeric x = X(:,j); % Convert x to array if x is a table. if istable(x) x = table2array(x); end % Group x into bins by using the discretize function. xbinned = discretize(x,[-inf; edges{j}; inf]); Xbinned(:,j) = xbinned; end
Xbinned contains the bin indices, ranging from 1 to the number of bins, for numeric predictors. Xbinned values are 0 for categorical predictors. If X contains NaNs, then the corresponding Xbinned values are NaNs.
Automatically optimize hyperparameters of an ECOC classifier by using fitcecoc.
Load the fisheriris data set.
load fisheriris
X = meas;
Y = species;Find hyperparameters that minimize the 5-fold cross-validation loss by using automatic hyperparameter optimization. For reproducibility, set the random seed and use the "expected-improvement-plus" acquisition function.
rng(0,"twister") hpoOptions = hyperparameterOptimizationOptions(AcquisitionFunctionName="expected-improvement-plus"); Mdl = fitcecoc(X,Y,OptimizeHyperparameters="auto", ... HyperparameterOptimizationOptions=hpoOptions)
|===================================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | Coding | BoxConstraint| KernelScale | Standardize |
| | result | | runtime | (observed) | (estim.) | | | | |
|===================================================================================================================================|
| 1 | Best | 0.3 | 9.2477 | 0.3 | 0.3 | onevsall | 76.389 | 0.0012205 | true |
| 2 | Best | 0.10667 | 0.15002 | 0.10667 | 0.1204 | onevsone | 0.0013787 | 41.108 | false |
| 3 | Best | 0.04 | 0.50859 | 0.04 | 0.135 | onevsall | 16.632 | 0.18987 | false |
| 4 | Accept | 0.046667 | 0.12768 | 0.04 | 0.079094 | onevsone | 0.04843 | 0.0042504 | true |
| 5 | Accept | 0.046667 | 0.66572 | 0.04 | 0.040197 | onevsall | 15.204 | 0.15933 | false |
| 6 | Accept | 0.08 | 0.069965 | 0.04 | 0.043201 | onevsall | 77.055 | 4.7599 | false |
| 7 | Accept | 0.16 | 5.0332 | 0.04 | 0.04347 | onevsall | 0.037396 | 0.0010042 | false |
| 8 | Accept | 0.046667 | 0.078502 | 0.04 | 0.043477 | onevsone | 0.0041486 | 0.32592 | true |
| 9 | Accept | 0.046667 | 0.10601 | 0.04 | 0.043118 | onevsone | 4.6545 | 0.041226 | true |
| 10 | Accept | 0.16667 | 0.069994 | 0.04 | 0.043001 | onevsone | 0.0030987 | 300.86 | true |
| 11 | Accept | 0.046667 | 2.0526 | 0.04 | 0.042997 | onevsone | 128.38 | 0.005555 | false |
| 12 | Accept | 0.046667 | 0.074196 | 0.04 | 0.043207 | onevsone | 0.081215 | 0.11353 | false |
| 13 | Accept | 0.33333 | 0.074429 | 0.04 | 0.043431 | onevsall | 243.89 | 987.69 | true |
| 14 | Accept | 0.14 | 2.0366 | 0.04 | 0.043265 | onevsone | 27.177 | 0.0010036 | true |
| 15 | Accept | 0.04 | 0.076787 | 0.04 | 0.040139 | onevsone | 0.0011464 | 0.001003 | false |
| 16 | Accept | 0.046667 | 0.089453 | 0.04 | 0.040165 | onevsone | 0.0010135 | 0.021485 | true |
| 17 | Accept | 0.046667 | 1.2196 | 0.04 | 0.040381 | onevsone | 0.42331 | 0.0010054 | false |
| 18 | Accept | 0.14 | 6.1481 | 0.04 | 0.04025 | onevsall | 956.72 | 0.053616 | false |
| 19 | Accept | 0.26667 | 0.070879 | 0.04 | 0.04023 | onevsall | 0.058487 | 1.2227 | false |
| 20 | Best | 0.04 | 0.072515 | 0.04 | 0.039873 | onevsone | 0.79359 | 1.4535 | true |
|===================================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | Coding | BoxConstraint| KernelScale | Standardize |
| | result | | runtime | (observed) | (estim.) | | | | |
|===================================================================================================================================|
| 21 | Accept | 0.04 | 0.076734 | 0.04 | 0.039837 | onevsone | 8.8581 | 1.123 | false |
| 22 | Accept | 0.04 | 0.069571 | 0.04 | 0.039802 | onevsone | 755.56 | 21.24 | false |
| 23 | Accept | 0.10667 | 0.076388 | 0.04 | 0.039824 | onevsone | 41.541 | 966.05 | false |
| 24 | Accept | 0.04 | 0.09998 | 0.04 | 0.039764 | onevsone | 966.21 | 0.33603 | false |
| 25 | Accept | 0.39333 | 7.5636 | 0.04 | 0.040001 | onevsall | 11.928 | 0.001094 | false |
| 26 | Accept | 0.04 | 0.075486 | 0.04 | 0.039986 | onevsone | 0.12946 | 0.092435 | true |
| 27 | Accept | 0.04 | 0.075959 | 0.04 | 0.039984 | onevsone | 6.8428 | 0.039038 | false |
| 28 | Accept | 0.04 | 0.07371 | 0.04 | 0.039983 | onevsone | 0.0010014 | 0.019004 | false |
| 29 | Accept | 0.04 | 0.068429 | 0.04 | 0.039985 | onevsone | 194.14 | 1.8004 | true |
| 30 | Accept | 0.046667 | 0.067375 | 0.04 | 0.039985 | onevsone | 769.43 | 141.77 | true |
__________________________________________________________
Optimization completed.
MaxObjectiveEvaluations of 30 reached.
Total function evaluations: 30
Total elapsed time: 43.9106 seconds
Total objective function evaluation time: 36.2198
Best observed feasible point:
Coding BoxConstraint KernelScale Standardize
________ _____________ ___________ ___________
onevsone 0.79359 1.4535 true
Observed objective function value = 0.04
Estimated objective function value = 0.040004
Function evaluation time = 0.072515
Best estimated feasible point (according to models):
Coding BoxConstraint KernelScale Standardize
________ _____________ ___________ ___________
onevsone 0.12946 0.092435 true
Estimated objective function value = 0.039985
Estimated function evaluation time = 0.07549

Mdl =
ClassificationECOC
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
BinaryLearners: {3×1 cell}
CodingName: 'onevsone'
HyperparameterOptimizationResults: [1×1 classreg.learning.paramoptim.SupervisedLearningBayesianOptimization]
Properties, Methods
The trained classifier Mdl corresponds to the best estimated feasible point and uses the same hyperparameter values for Coding, BoxConstraint, KernelScale, and Standardize.
Find the hyperparameter values used to train Mdl by using the bestPoint function. By default, bestPoint uses the same best point criterion used by fitcecoc during the hyperparameter optimization ("min-visited-upper-confidence-interval"). In general, fit functions determine the best hyperparameter values based on the "min-visited-upper-confidence-interval" criterion (instead of the "min-observed" criterion) to avoid overfitting to noise in the data set.
bestEstimatedPoint = bestPoint(Mdl.HyperparameterOptimizationResults)
bestEstimatedPoint=1×4 table
Coding BoxConstraint KernelScale Standardize
________ _____________ ___________ ___________
onevsone 0.12946 0.092435 true
Verify that the results match the properties of Mdl and its binary learners. Note that the template for the SVM binary learners contains a StandardizeData value instead of a Standardize value, but the two options are equivalent.
coding = Mdl.ModelParameters.Coding
coding = 'onevsone'
binaryLearnerProperties = Mdl.ModelParameters.BinaryLearners
binaryLearnerProperties =
Fit template for classification SVM.
Alpha: [0×1 double]
BoxConstraint: 0.1295
CacheSize: []
CachingMethod: ''
ClipAlphas: []
DeltaGradientTolerance: []
Epsilon: []
GapTolerance: []
KKTTolerance: []
IterationLimit: []
KernelFunction: ''
KernelScale: 0.0924
KernelOffset: []
KernelPolynomialOrder: []
NumPrint: []
Nu: []
OutlierFraction: []
RemoveDuplicates: []
ShrinkagePeriod: []
Solver: ''
StandardizeData: 1
SaveSupportVectors: 0
VerbosityLevel: []
Version: 2
Method: 'SVM'
Type: 'classification'
Create two multiclass ECOC models trained on tall data. Use linear binary learners for one of the models and kernel binary learners for the other. Compare the resubstitution classification error of the two models.
In general, you can perform multiclass classification of tall data by using fitcecoc with linear or kernel binary learners. When you use fitcecoc to train a model on tall arrays, you cannot use SVM binary learners directly. However, you can use either linear or kernel binary classification models that use SVMs.
When you perform calculations on tall arrays, MATLAB® uses either a parallel pool (default if you have Parallel Computing Toolbox™) or the local MATLAB session. If you want to run the example using the local MATLAB session when you have Parallel Computing Toolbox, you can change the global execution environment by using the mapreducer function.
Create a datastore that references the folder containing Fisher's iris data set. Specify 'NA' values as missing data so that datastore replaces them with NaN values. Create tall versions of the predictor and response data.
ds = datastore('fisheriris.csv','TreatAsMissing','NA'); t = tall(ds);
Starting parallel pool (parpool) using the 'local' profile ... Connected to the parallel pool (number of workers: 6).
X = [t.SepalLength t.SepalWidth t.PetalLength t.PetalWidth]; Y = t.Species;
Standardize the predictor data.
Z = zscore(X);
Train a multiclass ECOC model that uses tall data and linear binary learners. By default, when you pass tall arrays to fitcecoc, the software trains linear binary learners that use SVMs. Because the response data contains only three unique classes, change the coding scheme from one-versus-all (which is the default when you use tall data) to one-versus-one (which is the default when you use in-memory data).
For reproducibility, set the seeds of the random number generators using rng and tallrng. The results can vary depending on the number of workers and the execution environment for the tall arrays. For details, see Control Where Your Code Runs.
rng('default') tallrng('default') mdlLinear = fitcecoc(Z,Y,'Coding','onevsone')
Training binary learner 1 (Linear) out of 3. Training binary learner 2 (Linear) out of 3. Training binary learner 3 (Linear) out of 3.
mdlLinear =
CompactClassificationECOC
ResponseName: 'Y'
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
BinaryLearners: {3×1 cell}
CodingMatrix: [3×3 double]
Properties, Methods
mdlLinear is a CompactClassificationECOC model composed of three binary learners.
Train a multiclass ECOC model that uses tall data and kernel binary learners. First, create a templateKernel object to specify the properties of the kernel binary learners; in particular, increase the number of expansion dimensions to .
tKernel = templateKernel('NumExpansionDimensions',2^16)tKernel =
Fit template for classification Kernel.
BetaTolerance: []
BlockSize: []
BoxConstraint: []
Epsilon: []
NumExpansionDimensions: 65536
GradientTolerance: []
HessianHistorySize: []
IterationLimit: []
KernelScale: []
Lambda: []
Learner: 'svm'
LossFunction: []
Stream: []
VerbosityLevel: []
Version: 1
Method: 'Kernel'
Type: 'classification'
By default, the kernel binary learners use SVMs.
Pass the templateKernel object to fitcecoc and change the coding scheme to one-versus-one.
mdlKernel = fitcecoc(Z,Y,'Learners',tKernel,'Coding','onevsone')
Training binary learner 1 (Kernel) out of 3. Training binary learner 2 (Kernel) out of 3. Training binary learner 3 (Kernel) out of 3.
mdlKernel =
CompactClassificationECOC
ResponseName: 'Y'
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
BinaryLearners: {3×1 cell}
CodingMatrix: [3×3 double]
Properties, Methods
mdlKernel is also a CompactClassificationECOC model composed of three binary learners.
Compare the resubstitution classification error of the two models.
errorLinear = gather(loss(mdlLinear,Z,Y))
Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 1: Completed in 1.4 sec Evaluation completed in 1.6 sec
errorLinear = 0.0333
errorKernel = gather(loss(mdlKernel,Z,Y))
Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 1: Completed in 15 sec Evaluation completed in 16 sec
errorKernel = 0.0067
mdlKernel misclassifies a smaller percentage of the training data than mdlLinear.
Input Arguments
Sample data, specified as a table. Each row of Tbl corresponds
to one observation, and each column corresponds to one predictor.
Optionally, Tbl can contain one additional column
for the response variable. Multicolumn variables and cell arrays other
than cell arrays of character vectors are not accepted.
If Tbl contains the response variable, and
you want to use all remaining variables in Tbl as
predictors, then specify the response variable using ResponseVarName.
If Tbl contains the response variable, and
you want to use only a subset of the remaining variables in Tbl as
predictors, specify a formula using formula.
If Tbl does not contain the response variable,
specify a response variable using Y. The length
of response variable and the number of Tbl rows
must be equal.
Data Types: table
Response variable name, specified as the name of a variable in
Tbl.
You must specify ResponseVarName as a character vector or string scalar.
For example, if the response variable Y is
stored as Tbl.Y, then specify it as
"Y". Otherwise, the software
treats all columns of Tbl, including
Y, as predictors when training
the model.
The response variable must be a categorical, character, or string array; a logical or numeric
vector; or a cell array of character vectors. If
Y is a character array, then each
element of the response variable must correspond to one row of
the array.
A good practice is to specify the order of the classes by using the
ClassNames name-value
argument.
Data Types: char | string
Explanatory model of the response variable and a subset of the predictor variables,
specified as a character vector or string scalar in the form
"Y~x1+x2+x3". In this form, Y represents the
response variable, and x1, x2, and
x3 represent the predictor variables.
To specify a subset of variables in Tbl as predictors for
training the model, use a formula. If you specify a formula, then the software does not
use any variables in Tbl that do not appear in
formula.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl by
using the isvarname function. If the variable names
are not valid, then you can convert them by using the matlab.lang.makeValidName function.
Data Types: char | string
Class labels to which the ECOC model is trained, specified as a categorical, character, or string array, logical or numeric vector, or cell array of character vectors.
If Y is a character array, then each element
must correspond to one row of the array.
The length of Y and the number of rows of Tbl or X must
be equal.
It is good practice to specify the class order using the ClassNames name-value
pair argument.
Data Types: categorical | char | string | logical | single | double | cell
Predictor data, specified as a full or sparse matrix.
The length of Y and the number of observations
in X must be equal.
To specify the names of the predictors in the order of their
appearance in X, use the PredictorNames name-value
pair argument.
Note
For linear classification learners, if you orient
Xso that observations correspond to columns and specify'ObservationsIn','columns', then you can experience a significant reduction in optimization-execution time.For all other learners, orient
Xso that observations correspond to rows.fitcecocsupports sparse matrices for training linear classification models only.
Data Types: double | single
Note
The software treats NaN, empty character vector
(''), empty string (""),
<missing>, and <undefined>
elements as missing data. The software removes rows of X
corresponding to missing values in Y. However, the treatment of
missing values in X varies among binary learners. For details,
see the training functions for your binary learners: fitcdiscr, fitckernel, fitcknn, fitclinear, fitcnb, fitcsvm, fitctree, or fitcensemble. Removing observations decreases the effective training
or cross-validation sample size.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'Learners','tree','Coding','onevsone','CrossVal','on'
specifies to use decision trees for all binary learners, a one-versus-one coding
design, and to implement 10-fold cross-validation.
Note
You cannot use any cross-validation name-value argument together with the
OptimizeHyperparameters name-value argument. You can modify the
cross-validation for OptimizeHyperparameters only by using the
HyperparameterOptimizationOptions name-value argument.
ECOC Classifier Options
Coding design name, specified as the comma-separated pair consisting
of 'Coding' and a numeric matrix or a value in
this table.
| Value | Number of Binary Learners | Description |
|---|---|---|
"allpairs" and "onevsone" | K(K – 1)/2 | For each binary learner, one class is positive, another is negative, and the software ignores the rest. This design exhausts all combinations of class pair assignments. |
"binarycomplete" | This design partitions the classes into all binary combinations, and does not ignore any
classes. For each binary learner, all class assignments are
–1 and 1 with at least one positive
class and one negative class in the assignment. | |
"denserandom" | Random, but approximately 10 log2K | For each binary learner, the software randomly assigns classes into positive or negative classes, with at least one of each type. For more details, see Random Coding Design Matrices. |
"onevsall" | K | For each binary learner, one class is positive and the rest are negative. This design exhausts all combinations of positive class assignments. |
"ordinal" | K – 1 | For the first binary learner, the first class is negative and the rest are positive. For the second binary learner, the first two classes are negative and the rest are positive, and so on. |
"sparserandom" | Random, but approximately 15 log2K | For each binary learner, the software randomly assigns classes as positive or negative with probability 0.25 for each, and ignores classes with probability 0.5. For more details, see Random Coding Design Matrices. |
"ternarycomplete" | This design partitions the classes into all ternary combinations. All class assignments are
0, –1, and 1 with
at least one positive class and one negative class in each assignment. |
You can also specify a coding design using a custom coding matrix, which is a
K-by-L matrix. Each row corresponds to a class
and each column corresponds to a binary learner. The class order (rows) corresponds to
the order in ClassNames. Create the
matrix by following these guidelines:
Every element of the custom coding matrix must be
–1,0, or1, and the value must correspond to a dichotomous class assignment. ConsiderCoding(i,j), the class that learnerjassigns to observations in classi.Value Dichotomous Class Assignment –1Learner jassigns observations in classito a negative class.0Before training, learner jremoves observations in classifrom the data set.1Learner jassigns observations in classito a positive class.Every column must contain at least one
–1and one1.For all column indices
i,jwherei≠j,Coding(:,i)cannot equalCoding(:,j), andCoding(:,i)cannot equal–Coding(:,j).All rows of the custom coding matrix must be different.
For more details on the form of custom coding design matrices, see Custom Coding Design Matrices.
Example: 'Coding','ternarycomplete'
Data Types: char | string | double | single | int16 | int32 | int64 | int8
Flag indicating whether to transform scores to posterior probabilities,
specified as the comma-separated pair consisting of 'FitPosterior' and
a true (1) or false (0).
If FitPosterior is true,
then the software transforms binary-learner classification scores
to posterior probabilities. You can obtain posterior probabilities
by using kfoldPredict, predict,
or resubPredict.
fitcecoc does not support fitting posterior probabilities if:
The ensemble method is
AdaBoostM2,LPBoost,RUSBoost,RobustBoost, orTotalBoost.The binary learners (
Learners) are linear or kernel classification models that implement SVM. To obtain posterior probabilities for linear or kernel classification models, implement logistic regression instead.
Example: 'FitPosterior',true
Data Types: logical
Binary learner templates, specified as a character vector, string scalar, template object, or
cell vector of template objects. Specifically, you can specify binary classifiers such
as SVM, and the ensembles that use GentleBoost,
LogitBoost, and RobustBoost, to solve
multiclass problems. However, fitcecoc also supports multiclass
models as binary classifiers.
If
Learnersis a character vector or string scalar, then the software trains each binary learner using the default values of the specified algorithm. This table summarizes the available algorithms.Value Description "discriminant"Discriminant analysis. For default options, see templateDiscriminant."ensemble"(since R2024a)Ensemble learning model. By default, the ensemble uses an adaptive logistic regression ( "LogitBoost") aggregation method, 100 learning cycles, and tree weak learners. For other default options, seetemplateEnsemble."kernel"Kernel classification model. For default options, see templateKernel."knn"k-nearest neighbors. For default options, see templateKNN."linear"Linear classification model. For default options, see templateLinear."naivebayes"Naive Bayes. For default options, see templateNaiveBayes."svm"SVM. For default options, see templateSVM."tree"Classification trees. For default options, see templateTree.If
Learnersis a template object, then each binary learner trains according to the stored options. You can create a template object using:templateDiscriminant, for discriminant analysis.templateEnsemble, for ensemble learning. You must at least specify the learning method (Method), the number of learners (NLearn), and the type of learner (Learners). You cannot use the"AdaBoostM2"ensemble method for binary learning. If you want to perform hyperparameter optimization using theOptimizeHyperparametersname-value argument, the ensemble method must be"AdaBoostM1","GentleBoost", or"LogitBoost", and the ensemble weak learners must be trees.templateKernel, for kernel classification.templateKNN, for k-nearest neighbors.templateLinear, for linear classification.templateNaiveBayes, for naive Bayes.templateSVM, for SVM.templateTree, for classification trees.
If
Learnersis a cell vector of template objects, then:Cell j corresponds to binary learner j (in other words, column j of the coding design matrix), and the cell vector must have length L. L is the number of columns in the coding design matrix. For details, see
Coding.To use one of the built-in loss functions for prediction, then all binary learners must return a score in the same range. For example, you cannot include default SVM binary learners with default naive Bayes binary learners. The former returns a score in the range (-∞,∞), and the latter returns a posterior probability as a score. Otherwise, you must provide a custom loss as a function handle to functions such as
predictandloss.You cannot specify linear classification model learner templates with any other template.
Similarly, you cannot specify kernel classification model learner templates with any other template.
By default, the software trains learners using default SVM templates.
Example: "Learners","tree"
Number of bins for numeric predictors, specified as the
comma-separated pair consisting of 'NumBins' and a
positive integer scalar. This argument is valid only when
fitcecoc uses a tree learner, that is,
'Learners' is either 'tree'
or a template object created by using templateTree, or a template
object created by using templateEnsemble with tree
weak learners.
If the
NumBinsvalue is empty (default), thenfitcecocdoes not bin any predictors.If you specify the
NumBinsvalue as a positive integer scalar (numBins), thenfitcecocbins every numeric predictor into at mostnumBinsequiprobable bins, and then grows trees on the bin indices instead of the original data.The number of bins can be less than
numBinsif a predictor has fewer thannumBinsunique values.fitcecocdoes not bin categorical predictors.
When you use a large training data set, this binning option speeds up training but might cause
a potential decrease in accuracy. You can try NumBins=50 first, and then
change the value depending on the accuracy and training speed.
A trained model stores the bin edges in the BinEdges property.
Example: 'NumBins',50
Data Types: single | double
Number of binary learners concurrently trained, specified as the
comma-separated pair consisting of 'NumConcurrent'
and a positive integer scalar. The default value is
1, which means fitcecoc trains
the binary learners sequentially.
Note
This option applies only when you use
fitcecoc on tall arrays. See Tall Arrays
for more information.
Data Types: single | double
Predictor data observation dimension, specified as the comma-separated
pair consisting of 'ObservationsIn' and
'columns' or 'rows'.
Note
For linear classification learners, if you orient
Xso that observations correspond to columns and specify'ObservationsIn','columns', then you can experience a significant reduction in optimization-execution time.For all other learners, orient
Xso that observations correspond to rows.
Example: 'ObservationsIn','columns'
Verbosity level, specified as the comma-separated pair consisting of
'Verbose' and 0,
1, or 2.
Verbose controls the amount of diagnostic
information per binary learner that the software displays in the Command
Window.
This table summarizes the available verbosity level options.
| Value | Description |
|---|---|
0 | The software does not display diagnostic information. |
1 | The software displays diagnostic messages every time it trains a new binary learner. |
2 | The software displays extra diagnostic messages every time it trains a new binary learner. |
Each binary learner has its own verbosity level that is independent of
this name-value pair argument. To change the verbosity level of a binary
learner, create a template object and specify the
'Verbose' name-value pair argument. Then, pass
the template object to fitcecoc by using the
'Learners' name-value pair argument.
Example: 'Verbose',1
Data Types: double | single
Cross-Validation Options
Flag to train a cross-validated classifier, specified as the
comma-separated pair consisting of 'Crossval' and
'on' or 'off'.
If you specify 'on', then the software trains a
cross-validated classifier with 10 folds.
You can override this cross-validation setting using one of the
CVPartition, Holdout,
KFold, or Leaveout
name-value pair arguments. You can only use one cross-validation
name-value pair argument at a time to create a cross-validated
model.
Alternatively, cross-validate later by passing
Mdl to crossval.
Example: 'Crossval','on'
Cross-validation partition, specified as a cvpartition object that specifies the type of cross-validation and the
indexing for the training and validation sets.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Suppose you create a random partition for 5-fold cross-validation on 500
observations by using cvp = cvpartition(500,KFold=5). Then, you can
specify the cross-validation partition by setting
CVPartition=cvp.
Fraction of the data used for holdout validation, specified as a scalar value in the range
(0,1). If you specify Holdout=p, then the software completes these
steps:
Randomly select and reserve
p*100% of the data as validation data, and train the model using the rest of the data.Store the compact trained model in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Holdout=0.1
Data Types: double | single
Number of folds to use in the cross-validated model, specified as a positive integer value
greater than 1. If you specify KFold=k, then the software completes
these steps:
Randomly partition the data into
ksets.For each set, reserve the set as validation data, and train the model using the other
k– 1 sets.Store the
kcompact trained models in ak-by-1 cell vector in theTrainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: KFold=5
Data Types: single | double
Leave-one-out cross-validation flag, specified as the comma-separated
pair consisting of 'Leaveout' and
'on' or 'off'. If you specify
'Leaveout','on', then, for each of the
n observations, where n is
size(Mdl.X,1), the software:
Reserves the observation as validation data, and trains the model using the other n – 1 observations
Stores the n compact, trained models in the cells of a n-by-1 cell vector in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can use one of these four
options only: CVPartition,
Holdout, KFold, or
Leaveout.
Note
Leave-one-out is not recommended for cross-validating ECOC models composed of linear or kernel classification model learners.
Example: 'Leaveout','on'
Other Classification Options
Categorical predictors list, specified as one of the values in this table.
| Value | Description |
|---|---|
| Vector of positive integers |
Each entry in the vector is an index value indicating that the corresponding predictor is
categorical. The index values are between 1 and If |
| Logical vector |
A |
| Character matrix | Each row of the matrix is the name of a predictor variable. The names must match the entries in PredictorNames. Pad the names with extra blanks so each row of the character matrix has the same length. |
| String array or cell array of character vectors | Each element in the array is the name of a predictor variable. The names must match the entries in PredictorNames. |
"all" | All predictors are categorical. |
Specification of 'CategoricalPredictors' is
appropriate if:
At least one predictor is categorical and all binary learners are classification trees, naive Bayes learners, SVMs, linear learners, kernel learners, or ensembles of classification trees.
All predictors are categorical and at least one binary learner is kNN.
If you specify 'CategoricalPredictors'
for any other learner, then the software warns that it cannot train that
binary learner. For example, the software cannot train discriminant
analysis classifiers using categorical predictors.
Each learner identifies and treats categorical predictors in the same
way as the fitting function corresponding to the learner. See 'CategoricalPredictors' of
fitckernel for kernel learners, 'CategoricalPredictors' of fitcknn
for k-nearest learners, 'CategoricalPredictors' of
fitclinear for linear learners, 'CategoricalPredictors' of fitcnb
for naive Bayes learners, 'CategoricalPredictors' of fitcsvm
for SVM learners, and 'CategoricalPredictors' of fitctree
for tree learners.
Example: 'CategoricalPredictors','all'
Data Types: single | double | logical | char | string | cell
Names of classes to use for training, specified as a categorical, character, or string
array; a logical or numeric vector; or a cell array of character vectors.
ClassNames must have the same data type as the response variable
in Tbl or Y.
If ClassNames is a character array, then each element must correspond to one row of the array.
Use ClassNames to:
Specify the order of the classes during training.
Specify the order of any input or output argument dimension that corresponds to the class order. For example, use
ClassNamesto specify the order of the dimensions ofCostor the column order of classification scores returned bypredict.Select a subset of classes for training. For example, suppose that the set of all distinct class names in
Yis["a","b","c"]. To train the model using observations from classes"a"and"c"only, specifyClassNames=["a","c"].
The default value for ClassNames is the set of all distinct class names in the response variable in Tbl or Y.
Example: ClassNames=["b","g"]
Data Types: categorical | char | string | logical | single | double | cell
Misclassification cost, specified as the comma-separated pair
consisting of 'Cost' and a square matrix or
structure. If you specify:
The square matrix
Cost, thenCost(i,j)is the cost of classifying a point into classjif its true class isi. That is, the rows correspond to the true class and the columns correspond to the predicted class. To specify the class order for the corresponding rows and columns ofCost, additionally specify theClassNamesname-value pair argument.The structure
S, then it must have two fields:S.ClassNames, which contains the class names as a variable of the same data type asYS.ClassificationCosts, which contains the cost matrix with rows and columns ordered as inS.ClassNames
The default is ones(, where
K) -
eye(K)K is the number of distinct
classes.
Example: 'Cost',[0 1 2 ; 1 0 2; 2 2
0]
Data Types: double | single | struct
Parallel computing options, specified as the comma-separated pair consisting of
'Options' and a structure array returned by statset. Parallel computation requires Parallel Computing
Toolbox™. fitcecoc uses
'Streams', 'UseParallel', and
'UseSubtreams' fields.
This table summarizes the available options.
| Option | Description |
|---|---|
'Streams' | A
In that case, use a cell array of
the same size as the parallel pool. If a parallel pool is not
open, then the software tries to open one (depending on your
settings), and |
'UseParallel' | If you have Parallel Computing Toolbox, then you can invoke a
pool of workers by setting
When you use
decision trees for binary learners,
|
'UseSubstreams' | Set to true to compute using the stream specified by
'Streams'. Default is
false. For example, set
Streams to a type allowing substreams, such
as'mlfg6331_64' or
'mrg32k3a'. |
A best practice to ensure more
predictable results is to use parpool (Parallel Computing Toolbox) and
explicitly create a parallel pool before you invoke parallel computing
using fitcecoc.
Example: 'Options',statset('UseParallel',true)
Data Types: struct
Predictor variable names, specified as a string array of unique names or cell array of unique
character vectors. The functionality of PredictorNames depends on the
way you supply the training data.
If you supply
XandY, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the column order ofX. That is,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'x1','x2',...}.
If you supply
Tbl, then you can usePredictorNamesto choose which predictor variables to use in training. That is,fitcecocuses only the predictor variables inPredictorNamesand the response variable during training.PredictorNamesmust be a subset ofTbl.Properties.VariableNamesand cannot include the name of the response variable.By default,
PredictorNamescontains the names of all predictor variables.A good practice is to specify the predictors for training using either
PredictorNamesorformula, but not both.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Prior probabilities for each class, specified as the comma-separated
pair consisting of 'Prior' and a value in this
table.
| Value | Description |
|---|---|
'empirical' | The class prior probabilities are the class
relative frequencies in
Y. |
'uniform' | All class prior probabilities are equal to 1/K, where K is the number of classes. |
| numeric vector | Each element is a class prior probability. Order
the elements according to
Mdl.ClassNames
or specify the order using the
ClassNames name-value pair
argument. The software normalizes the elements such
that they sum to 1. |
| structure |
A structure
|
For more details on how the software incorporates class prior probabilities, see Prior Probabilities and Misclassification Cost.
Example: struct('ClassNames',{{'setosa','versicolor','virginica'}},'ClassProbs',1:3)
Data Types: single | double | char | string | struct
Response variable name, specified as a character vector or string scalar.
If you supply
Y, then you can useResponseNameto specify a name for the response variable.If you supply
ResponseVarNameorformula, then you cannot useResponseName.
Example: ResponseName="response"
Data Types: char | string
Observation weights, specified as a numeric vector of positive values or name of a variable in
Tbl. The software weighs the observations in each row of
X or Tbl with the corresponding value in
Weights. The size of Weights must equal the
number of rows of X or Tbl.
If you specify the input data as a table Tbl, then
Weights can be the name of a variable in Tbl
that contains a numeric vector. In this case, you must specify
Weights as a character vector or string scalar. For example, if
the weights vector W is stored as Tbl.W, then
specify it as "W". Otherwise, the software treats all columns of
Tbl, including W, as predictors or the
response when training the model.
By default, Weights is
ones(, where
n,1)n is the number of observations in X
or Tbl.
The software normalizes Weights to sum up to the value of the prior
probability in the respective class. Inf weights are not supported.
Data Types: double | single | char | string
Hyperparameter Optimization
Parameters to optimize, specified as one of the following:
"none"— Do not optimize."auto"— Use"Coding"along with the default parameters for the specifiedLearners:Learners="svm"(default) —["BoxConstraint","KernelScale","Standardize"]Learners="discriminant"—["Delta","Gamma"]Learners="ensemble"—["LearnRate","Method","MinLeafSize","NumLearningCycles"]Learners="kernel"—["KernelScale","Lambda","Standardize"]Learners="knn"—["Distance","NumNeighbors","Standardize"]Learners="linear"—["Lambda","Learner"]Learners="tree"—"MinLeafSize"
"all"— Optimize all eligible parameters.String array or cell array of eligible parameter names
Vector of
optimizableVariableobjects, typically the output ofhyperparameters
The optimization attempts to minimize the cross-validation loss
(error) for fitcecoc by varying the parameters. To control the
cross-validation type and other aspects of the optimization, use the
HyperparameterOptimizationOptions name-value argument. When you use
HyperparameterOptimizationOptions, you can use the (compact) model size
instead of the cross-validation loss as the optimization objective by setting the
ConstraintType and ConstraintBounds options.
Note
The values of OptimizeHyperparameters override any values you
specify using other name-value arguments. For example, setting
OptimizeHyperparameters to "auto" causes
fitcecoc to optimize hyperparameters corresponding to the
"auto" option and to ignore any specified values for the
hyperparameters.
The eligible parameters for fitcecoc are:
Coding—fitcecocsearches among"onevsall"and"onevsone".The eligible hyperparameters for the chosen
Learners, as specified in this table.Learners Eligible Hyperparameters
(Bold = Default)Default Range "discriminant"DeltaLog-scaled in the range [1e-6,1e3]GammaReal values in [0,1]DiscrimType"linear","quadratic","diagLinear","diagQuadratic","pseudoLinear", and"pseudoQuadratic""ensemble"(since R2024a)Method"AdaBoostM1","GentleBoost", and"LogitBoost"NumLearningCyclesIntegers log-scaled in the range [10,500]LearnRatePositive values log-scaled in the range [1e-3,1]MinLeafSizeIntegers log-scaled in the range [1,max(2,floor(NumObservations/2))]MaxNumSplitsIntegers log-scaled in the range [1,max(2,NumObservations-1)]SplitCriterion"deviance","gdi", and"twoing""kernel"Learner"svm"and"logistic"KernelScalePositive values log-scaled in the range [1e-3,1e3]LambdaPositive values log-scaled in the range [1e-3/NumObservations,1e3/NumObservations]NumExpansionDimensionsIntegers log-scaled in the range [100,10000]Standardize"true"and"false""knn"NumNeighborsPositive integer values log-scaled in the range [1, max(2,round(NumObservations/2))]Distance"cityblock","chebychev","correlation","cosine","euclidean","hamming","jaccard","mahalanobis","minkowski","seuclidean", and"spearman"DistanceWeight"equal","inverse", and"squaredinverse"ExponentPositive values in [0.5,3]Standardize"true"and"false""linear"LambdaPositive values log-scaled in the range [1e-5/NumObservations,1e5/NumObservations]Learner"svm"and"logistic"Regularization"ridge"and"lasso"When
Regularizationis"ridge", the function uses a Limited-memory BFGS (LBFGS) solver by default.When
Regularizationis"lasso", the function uses a Sparse Reconstruction by Separable Approximation (SpaRSA) solver by default.
"svm"BoxConstraintPositive values log-scaled in the range [1e-3,1e3]KernelScalePositive values log-scaled in the range [1e-3,1e3]KernelFunction"gaussian","linear", and"polynomial"PolynomialOrderIntegers in the range [2,4]Standardize"true"and"false""tree"MinLeafSizeIntegers log-scaled in the range [1,max(2,floor(NumObservations/2))]MaxNumSplitsIntegers log-scaled in the range [1,max(2,NumObservations-1)]SplitCriterion"gdi","deviance", and"twoing"NumVariablesToSampleIntegers in the range [1,max(2,NumPredictors)]Alternatively, use
hyperparameterswith your chosenLearners, such asload fisheriris % hyperparameters requires data and learner params = hyperparameters("fitcecoc",meas,species,"svm");
To see the eligible and default hyperparameters, examine
params.
Set nondefault parameters by passing a vector of
optimizableVariable objects that have nondefault
values. For example,
load fisheriris params = hyperparameters("fitcecoc",meas,species,"svm"); params(2).Range = [1e-4,1e6];
Pass params as the value of
OptimizeHyperparameters.
By default, the iterative display appears at the command line,
and plots appear according to the number of hyperparameters in the optimization. For the
optimization and plots, the objective function is the misclassification rate. To control the
iterative display, set the Verbose option of the
HyperparameterOptimizationOptions name-value argument. To control the
plots, set the ShowPlots field of the
HyperparameterOptimizationOptions name-value argument.
For an example, see Optimize ECOC Classifier.
Example: "OptimizeHyperparameters","auto"
Options for optimization, specified as a HyperparameterOptimizationOptions object or a structure.
This argument modifies the effect of the
OptimizeHyperparameters name-value argument. If
you specify HyperparameterOptimizationOptions, you
must also specify OptimizeHyperparameters. All the
options are optional. However, you must set
ConstraintBounds and
ConstraintType to return
AggregateOptimizationResults. The options that
you can set in a structure are the same as those in the
HyperparameterOptimizationOptions object.
| Option | Values | Default |
|---|---|---|
Optimizer |
| "bayesopt" |
ConstraintBounds | Constraint bounds for N
optimization problems, specified as an
N-by-2 numeric matrix or
| [] |
ConstraintTarget | Constraint target for the optimization
problems, specified as | If you specify ConstraintBounds
and ConstraintType, then the default
value is "matlab". Otherwise, the
default value is []. |
ConstraintType | Constraint type for the optimization problems,
specified as | [] |
AcquisitionFunctionName | Type of acquisition function:
Acquisition
functions whose names include
| "expected-improvement-per-second-plus" |
LossFun | Type of validation loss to optimize, specified as
"auto",
"classifcost", or
"classiferror". In the case of
fitcecoc, the
"auto" and
"classiferror" options are
equivalent, and the software uses the misclassified rate
in decimal. "classifcost" indicates
to use the observed misclassification cost. | "auto" |
MaxObjectiveEvaluations | Maximum number of objective function evaluations. If
you specify multiple optimization problems using
ConstraintBounds, the value of
MaxObjectiveEvaluations applies
to each optimization problem individually. |
|
MaxTime | Time limit for the optimization, specified as a
nonnegative real scalar. The time limit is in
seconds, as measured by | Inf |
NumGridDivisions | For Optimizer="gridsearch", the
number of values in each dimension. The value can be a
vector of positive integers giving the number of values
for each dimension, or a scalar that applies to all
dimensions. This option is ignored for categorical
variables. | 10 |
ShowPlots | Logical value indicating whether to show plots of the
optimization progress. If this option is
true, the software plots the best
observed objective function value against the iteration
number. If you use Bayesian optimization
(Optimizer="bayesopt"),
then this field also plots the best estimated objective
function value. The best observed objective function
values and best estimated objective function values
correspond to the values in the BestSoFar
(observed) and BestSoFar
(estim.) columns of the iterative display,
respectively. You can find these values in the
properties ObjectiveMinimumTrace and EstimatedObjectiveMinimumTrace of the
SupervisedLearningBayesianOptimization
object. If the problem includes one or two optimization
parameters for Bayesian optimization, then
ShowPlots also plots a model of
the objective function against the parameters. | true |
SaveIntermediateResults | Logical value indicating whether to save the
optimization results. If this option is
true, the software overwrites a
workspace variable named
SupervisedLearningBayesoptResults
at each iteration. The variable is a SupervisedLearningBayesianOptimization
object. If you specify multiple optimization problems
using ConstraintBounds, the
workspace variable is an AggregateBayesianOptimization object named
AggregateBayesoptResults. | false |
Verbose | Display level at the command line:
For details, see the
| 1 |
UseParallel | Logical value indicating whether to run the Bayesian optimization in parallel, which requires Parallel Computing Toolbox. Due to the nonreproducibility of parallel timing, parallel Bayesian optimization does not necessarily yield reproducible results. For details, see Parallel Bayesian Optimization. | false |
Repartition | Logical value indicating whether to repartition
the cross-validation at every iteration. If this
option is A value of
| false |
| Specify only one of the following three options. | ||
CVPartition | cvpartition object created by
cvpartition | Kfold=5 if you do not
specify a cross-validation option |
Holdout | Scalar in the range (0,1)
representing the holdout fraction | |
Kfold | Integer greater than 1 | |
Example: HyperparameterOptimizationOptions=struct(UseParallel=true)
Output Arguments
Trained ECOC classifier, returned as a ClassificationECOC object, a
CompactClassificationECOC object,
a ClassificationPartitionedECOC
object, a ClassificationPartitionedKernelECOC object, a ClassificationPartitionedLinearECOC object, or a cell array of
model objects.
If you set any of the name-value arguments
CrossVal,CVPartition,Holdout,KFold, orLeaveout, thenMdlis aClassificationPartitionedECOC,ClassificationPartitionedKernelECOC, orClassificationPartitionedLinearECOCobject, depending on theLearnersname-value argument.If you specify
OptimizeHyperparametersand set theConstraintTypeandConstraintBoundsoptions ofHyperparameterOptimizationOptions, thenMdlis an N-by-1 cell array of model objects, where N is equal to the number of rows inConstraintBounds. If none of the optimization problems yields a feasible model, then each cell array value is[].Otherwise,
Mdlis aClassificationECOCorCompactClassificationECOCmodel object, depending on theLearnersname-value argument. That is,Mdlis a compact ECOC model if you use kernel or linear binary learners, and a full ECOC model otherwise.
To reference properties of a model object, use dot notation.
Cross-validation optimization of hyperparameters, returned as a SupervisedLearningBayesianOptimization object, an AggregateBayesianOptimization object, or a table of
hyperparameters and associated values.
HyperparameterOptimizationResults is nonempty when
the OptimizeHyperparameters name-value argument is
nonempty and either of the following conditions applies:
The
Learnersvalue is"linear"or"kernel".The
ConstraintTypeandConstraintBoundsoptions of theHyperparameterOptimizationOptionsname-value argument are nonempty.
If you specify ConstraintType and
ConstraintBounds, then
HyperparameterOptimizationResults is an
AggregateBayesianOptimization object. Otherwise, the
value of HyperparameterOptimizationResults depends on
the value of the Optimizer option in
HyperparameterOptimizationOptions.
Value of Optimizer Option | Value of HyperparameterOptimizationResults |
|---|---|
"bayesopt" (default) | SupervisedLearningBayesianOptimization object |
"gridsearch" or "randomsearch" | Table of hyperparameters used, observed objective function values (cross-validation loss), and observation ranks from lowest (best) to highest (worst) |
Limitations
fitcecocsupports sparse matrices for training linear classification models only. For all other models, supply a full matrix of predictor data instead.
More About
An error-correcting output codes (ECOC) model reduces the problem of classification with three or more classes to a set of binary classification problems.
ECOC classification requires a coding design, which determines the classes that the binary learners train on, and a decoding scheme, which determines how the results (predictions) of the binary classifiers are aggregated.
Assume the following:
The classification problem has three classes.
The coding design is one-versus-one. For three classes, this coding design is
You can specify a different coding design by using the
Codingname-value argument when you create a classification model.The model determines the predicted class by using the loss-weighted decoding scheme with the binary loss function g. The software also supports the loss-based decoding scheme. You can specify the decoding scheme and binary loss function by using the
DecodingandBinaryLossname-value arguments, respectively, when you call object functions, such aspredict,loss,margin,edge, and so on.
The ECOC algorithm follows these steps.
Learner 1 trains on observations in Class 1 or Class 2, and treats Class 1 as the positive class and Class 2 as the negative class. The other learners are trained similarly.
Let M be the coding design matrix with elements mkl, and sl be the predicted classification score for the positive class of learner l. The algorithm assigns a new observation to the class () that minimizes the aggregation of the losses for the B binary learners.
ECOC models can improve classification accuracy, compared to other multiclass models [2].
The coding design is a matrix whose elements direct which classes are trained by each binary learner, that is, how the multiclass problem is reduced to a series of binary problems.
Each row of the coding design corresponds to a distinct class, and each column corresponds to a binary learner. In a ternary coding design, for a particular column (or binary learner):
A row containing 1 directs the binary learner to group all observations in the corresponding class into a positive class.
A row containing –1 directs the binary learner to group all observations in the corresponding class into a negative class.
A row containing 0 directs the binary learner to ignore all observations in the corresponding class.
Coding design matrices with large, minimal, pairwise row distances based on the Hamming measure are optimal. For details on the pairwise row distance, see Random Coding Design Matrices and [3].
This table describes popular coding designs.
| Coding Design | Description | Number of Learners | Minimal Pairwise Row Distance |
|---|---|---|---|
| one-versus-all (OVA) | For each binary learner, one class is positive and the rest are negative. This design exhausts all combinations of positive class assignments. | K | 2 |
| one-versus-one (OVO) | For each binary learner, one class is positive, one class is negative, and the rest are ignored. This design exhausts all combinations of class pair assignments. | K(K – 1)/2 | 1 |
| binary complete | This design partitions the classes into all binary
combinations, and does not ignore any classes. That is, all class
assignments are | 2K – 1 – 1 | 2K – 2 |
| ternary complete | This design partitions the classes into all ternary
combinations. That is, all class assignments are
| (3K – 2K + 1 + 1)/2 | 3K – 2 |
| ordinal | For the first binary learner, the first class is negative and the rest are positive. For the second binary learner, the first two classes are negative and the rest are positive, and so on. | K – 1 | 1 |
| dense random | For each binary learner, the software randomly assigns classes into positive or negative classes, with at least one of each type. For more details, see Random Coding Design Matrices. | Random, but approximately 10 log2K | Variable |
| sparse random | For each binary learner, the software randomly assigns classes as positive or negative with probability 0.25 for each, and ignores classes with probability 0.5. For more details, see Random Coding Design Matrices. | Random, but approximately 15 log2K | Variable |
This plot compares the number of binary learners for the coding designs with an increasing number of classes (K).
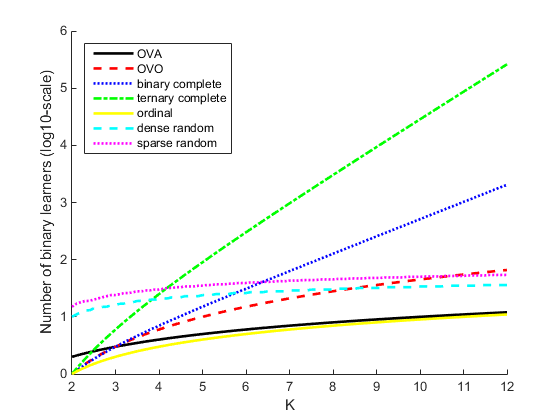
Tips
The number of binary learners grows with the number of classes. For a problem with many classes, the
binarycompleteandternarycompletecoding designs are not efficient. However:If K ≤ 4, then use
ternarycompletecoding design rather thansparserandom.If K ≤ 5, then use
binarycompletecoding design rather thandenserandom.
You can display the coding design matrix of a trained ECOC classifier by entering
Mdl.CodingMatrixinto the Command Window.You should form a coding matrix using intimate knowledge of the application, and taking into account computational constraints. If you have sufficient computational power and time, then try several coding matrices and choose the one with the best performance (e.g., check the confusion matrices for each model using
confusionchart).Leave-one-out cross-validation (
Leaveout) is inefficient for data sets with many observations. Instead, use k-fold cross-validation (KFold).
After training a model, you can generate C/C++ code that predicts labels for new data. Generating C/C++ code requires MATLAB Coder™. For details, see Introduction to Code Generation for Statistics and Machine Learning Functions.
Algorithms
Custom coding matrices must have a certain form. The software validates a custom coding matrix by ensuring:
Every element is –1, 0, or 1.
Every column contains as least one –1 and one 1.
For all distinct column vectors u and v, u ≠ v and u ≠ –v.
All row vectors are unique.
The matrix can separate any two classes. That is, you can move from any row to any other row following these rules:
Move vertically from 1 to –1 or –1 to 1.
Move horizontally from a nonzero element to another nonzero element.
Use a column of the matrix for a vertical move only once.
If it is not possible to move from row i to row j using these rules, then classes i and j cannot be separated by the design. For example, in the coding design
classes 1 and 2 cannot be separated from classes 3 and 4 (that is, you cannot move horizontally from –1 in row 2 to column 2 because that position contains a 0). Therefore, the software rejects this coding design.
If you use parallel computing (see Options),
then fitcecoc trains binary learners in parallel.
If you specify the Cost,
Prior, and Weights name-value arguments, the
output model object stores the specified values in the Cost,
Prior, and W properties, respectively. The
Cost property stores the user-specified cost matrix as is. The
Prior and W properties store the prior probabilities
and observation weights, respectively, after normalization. For details, see Misclassification Cost Matrix, Prior Probabilities, and Observation Weights.
For each binary learner, the software normalizes the prior probabilities into a
vector of two elements, and normalizes the cost matrix into a 2-by-2 matrix. Then,
the software adjusts the prior probability vector by incorporating the penalties
described in the 2-by-2 cost matrix, and sets the cost matrix to the default cost
matrix. The Cost and Prior properties of the
binary learners in Mdl (Mdl.BinaryLearners)
store the adjusted values. Specifically, the software completes these steps:
The software normalizes the specified class prior probabilities (
Prior) for each binary learner. Let M be the coding design matrix and I(A,c) be an indicator matrix. The indicator matrix has the same dimensions as A. If the corresponding element of A is c, then the indicator matrix has elements equaling one, and zero otherwise. Let M+1 and M-1 be K-by-L matrices such that:M+1 = M○I(M,1), where ○ is element-wise multiplication (that is,
Mplus = M.*(M == 1)). Also, let be column vector l of M+1.M-1 = -M○I(M,-1) (that is,
Mminus = -M.*(M == -1)). Also, let be column vector l of M-1.
Let and , where π is the vector of specified, class prior probabilities (
Prior).Then, the positive and negative, scalar class prior probabilities for binary learner l are
where j = {-1,1} and is the one-norm of a.
The software normalizes the K-by-K cost matrix C (
Cost) for each binary learner. For binary learner l, the cost of classifying a negative-class observation into the positive class isSimilarly, the cost of classifying a positive-class observation into the negative class is
The cost matrix for binary learner l is
ECOC models accommodate misclassification costs by incorporating them with class prior probabilities. The software adjusts the class prior probabilities and sets the cost matrix to the default cost matrix for binary learners as follows:
For a given number of classes K, the software generates random coding design matrices as follows.
The software generates one of these matrices:
Dense random — The software assigns 1 or –1 with equal probability to each element of the K-by-Ld coding design matrix, where .
Sparse random — The software assigns 1 to each element of the K-by-Ls coding design matrix with probability 0.25, –1 with probability 0.25, and 0 with probability 0.5, where .
If a column does not contain at least one 1 and one –1, then the software removes that column.
For distinct columns u and v, if u = v or u = –v, then the software removes v from the coding design matrix.
The software randomly generates 10,000 matrices by default, and retains the matrix with the largest, minimal, pairwise row distance based on the Hamming measure ([3]) given by
where mkjl is an element of coding design matrix j.
By default and for efficiency, fitcecoc empties the Alpha, SupportVectorLabels,
and SupportVectors properties
for all linear SVM binary learners. fitcecoc lists Beta, rather than
Alpha, in the model display.
To store Alpha, SupportVectorLabels, and
SupportVectors, pass a linear SVM template that specifies storing
support vectors to fitcecoc. For example,
enter:
t = templateSVM('SaveSupportVectors',true) Mdl = fitcecoc(X,Y,'Learners',t);
You can remove the support vectors and related values by passing the resulting
ClassificationECOC model to
discardSupportVectors.
References
[1] Allwein, E., R. Schapire, and Y. Singer. “Reducing multiclass to binary: A unifying approach for margin classifiers.” Journal of Machine Learning Research. Vol. 1, 2000, pp. 113–141.
[2] Fürnkranz, Johannes. “Round Robin Classification.” J. Mach. Learn. Res., Vol. 2, 2002, pp. 721–747.
[3] Escalera, S., O. Pujol, and P. Radeva. “Separability of ternary codes for sparse designs of error-correcting output codes.” Pattern Recog. Lett. Vol. 30, Issue 3, 2009, pp. 285–297.
[4] Escalera, S., O. Pujol, and P. Radeva. “On the decoding process in ternary error-correcting output codes.” IEEE Transactions on Pattern Analysis and Machine Intelligence. Vol. 32, Issue 7, 2010, pp. 120–134.
Extended Capabilities
The
fitcecoc function supports tall arrays with the following usage
notes and limitations:
Supported syntaxes are:
Mdl = fitcecoc(X,Y)Mdl = fitcecoc(X,Y,Name,Value)[Mdl,FitInfo,HyperparameterOptimizationResults] = fitcecoc(X,Y,Name,Value)—fitcecocreturns the additional output argumentsFitInfoandHyperparameterOptimizationResultswhen you specify the'OptimizeHyperparameters'name-value pair argument.
The
FitInfooutput argument is an empty structure array currently reserved for possible future use.Options related to cross-validation are not supported. The supported name-value pair arguments are:
'ClassNames''Cost''Coding'— Default value is'onevsall'.HyperparameterOptimizationOptions— For cross-validation, tall optimization supports onlyHoldoutvalidation. By default, the software selects and reserves 20% of the data as holdout validation data, and trains the model using the rest of the data. You can specify a different value for the holdout fraction by using this argument. For example, specifyHyperparameterOptimizationOptions=struct(Holdout=0.3)to reserve 30% of the data as validation data.'Learners'— Default value is'linear'. You can specify'linear','kernel', atemplateLinearortemplateKernelobject, or a cell array of such objects.'OptimizeHyperparameters'— When you use linear binary learners, the value of the'Regularization'hyperparameter must be'ridge'.'Prior''Verbose'— Default value is1.'Weights'
This additional name-value pair argument is specific to tall arrays:
'NumConcurrent'— A positive integer scalar specifying the number of binary learners that are trained concurrently by combining file I/O operations. The default value for'NumConcurrent'is1, which meansfitcecoctrains the binary learners sequentially.'NumConcurrent'is most beneficial when the input arrays cannot fit into the distributed cluster memory. Otherwise, the input arrays can be cached and speedup is negligible.If you run your code on Apache® Spark™,
NumConcurrentis upper bounded by the memory available for communications. Check the'spark.executor.memory'and'spark.driver.memory'properties in your Apache Spark configuration. Seeparallel.cluster.Hadoop(Parallel Computing Toolbox) for more details. For more information on Apache Spark and other execution environments that control where your code runs, see Extend Tall Arrays with Other Products.
For more information, see Tall Arrays.
To run in parallel, set the 'UseParallel' option to
true in one of these ways:
Set the
'UseParallel'option of the options structure totrueusingstatsetand specify the'Options'name-value pair argument in the call tofitceoc.For example:
'Options',statset('UseParallel',true)For more information, see the
'Options'name-value pair argument.Perform parallel hyperparameter optimization by using the
UseParallel=trueoption in theHyperparameterOptimizationOptionsname-value pair argument in the call tofitceoc.For more information on parallel hyperparameter optimization, see Parallel Bayesian Optimization.
For general information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Usage notes and limitations:
You can specify the name-value argument
Learnersonly as one of the learners specified in this table.Learner Template Object Creation Function gpuArraySupport"ensemble"templateEnsembleGPU Arrays for fitcensemble"kernel"templateKernelGPU Arrays for fitckernel"knn"templateKNNGPU Arrays for fitcknn"linear"templateLinearGPU Arrays for fitclinear"svm"templateSVMGPU Arrays for fitcsvm"tree"templateTreeGPU Arrays for fitctree
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2014bYou can optimize classification model hyperparameters with respect to
misclassification cost. When you use a classification fit function, specify the
OptimizeHyperparameters and
HyperparameterOptimizationOptions name-value arguments. In
the HyperparameterOptimizationOptions structure or object, set
the LossFun value to "classifcost".
Depending on the type of supervised learning fit function you use to perform
hyperparameter optimization, you can set the LossFun value to
"classifcost", "classiferror",
"mincost", "mse", or
"quantile".
If you perform Bayesian hyperparameter optimization using
fitcecoc with linear or kernel binary learners, you can
return the optimization results
(HyperparameterOptimizationResults), which are stored in a
SupervisedLearningBayesianOptimization object. In previous releases,
the optimization results are stored in a BayesianOptimization object.
You can specify kernel learners when you pass gpuArray inputs to
fitcecoc.
fitcecoc defaults to serial hyperparameter optimization when
HyperparameterOptimizationOptions includes
UseParallel=true and the software cannot open a parallel pool.
In previous releases, the software issues an error under these circumstances.
You can now specify linear and ensemble learners when you pass
gpuArray inputs to fitcecoc.
fitcecoc supports hyperparameter
optimization when you use ensemble binary learners. You can specify the OptimizeHyperparameters name-value argument when the Learners value is
"ensemble" or an ensemble template created using the
templateEnsemble function.
When you perform hyperparameter optimization using ensemble binary learners, the ensemble method must be
"AdaBoostM1","GentleBoost", or"LogitBoost", and the ensemble weak learners must be trees.The
"ensemble"value corresponds to an ensemble that uses an adaptive logistic regression aggregation method (LogitBoost), 100 learning cycles, and tree weak learners.
fitcecoc uses a default value of 70 for
MaxObjectiveEvaluations when performing Bayesian optimization
with ensemble binary learners. For more information, see HyperparameterOptimizationOptions.
You can use the hyperparameters function to see the
eligible and default hyperparameters for the ensemble binary learners. Additionally,
you can use the function to adjust the hyperparameters and their ranges. Specify
"fitcecoc" as the fitting function name and
"ensemble" or an ensemble template as the learner
type.
Starting in R2023b, when you specify "kernel",
"knn", or "svm" as the
Learners value and "auto" as the
OptimizeHyperparameters value,
fitcecoc includes Standardize as an
optimizable hyperparameter.
Starting in R2022a, when you specify to optimize hyperparameters for an ECOC model
with linear binary learners ('linear' or templateLinear) and do not specify to use a particular solver,
fitcecoc uses either a Limited-memory BFGS (LBFGS)
solver or a Sparse Reconstruction by Separable Approximation (SpaRSA) solver,
depending on the regularization type selected during each iteration of the
hyperparameter optimization.
When
Regularizationis'ridge', the function sets theSolvervalue to'lbfgs'by default.When
Regularizationis'lasso', the function sets theSolvervalue to'sparsa'by default.
In previous releases, the default solver selection during hyperparameter
optimization depended on various factors, including the regularization type, learner
type, and number of predictors. For more information, see Solver.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
选择网站
选择网站以获取翻译的可用内容,以及查看当地活动和优惠。根据您的位置,我们建议您选择:。
您也可以从以下列表中选择网站:
如何获得最佳网站性能
选择中国网站(中文或英文)以获得最佳网站性能。其他 MathWorks 国家/地区网站并未针对您所在位置的访问进行优化。
美洲
- América Latina (Español)
- Canada (English)
- United States (English)
欧洲
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)